dfvue is a minimal GUI for a quick view of csv files. It uses an
input panel similar to Microsoft Excel to check visually that the csv
file is read correctly. It provides most options of pandas' read_csv
method to be very versatile on the possible csv format.
dfvue is a Python script that can be called from within Python or
as a command line tool. It is not supposed to produce
publication-ready plots but rather provide a quick overview of the csv
file.
The complete documentation for dfvue is available from:
https://mcuntz.github.io/dfvue/
dfvue can be run from the command line:
dfvue csv_file.csvor from within Python:
from dfvue import dfvue
dfvue('csv_file.csv')where the csv file is optional. The latter can be left out and a csv
file can be opened with the "Open File" button from within dfvue.
Note, dfvue uses the TkAgg backend of matplotlib. It must be
called before any other call to matplotlib. This also means that you
cannot launch it from within iPython if it was launched with
--pylab. It can be called from within a standard iPython, though,
or using ipython --gui tk.
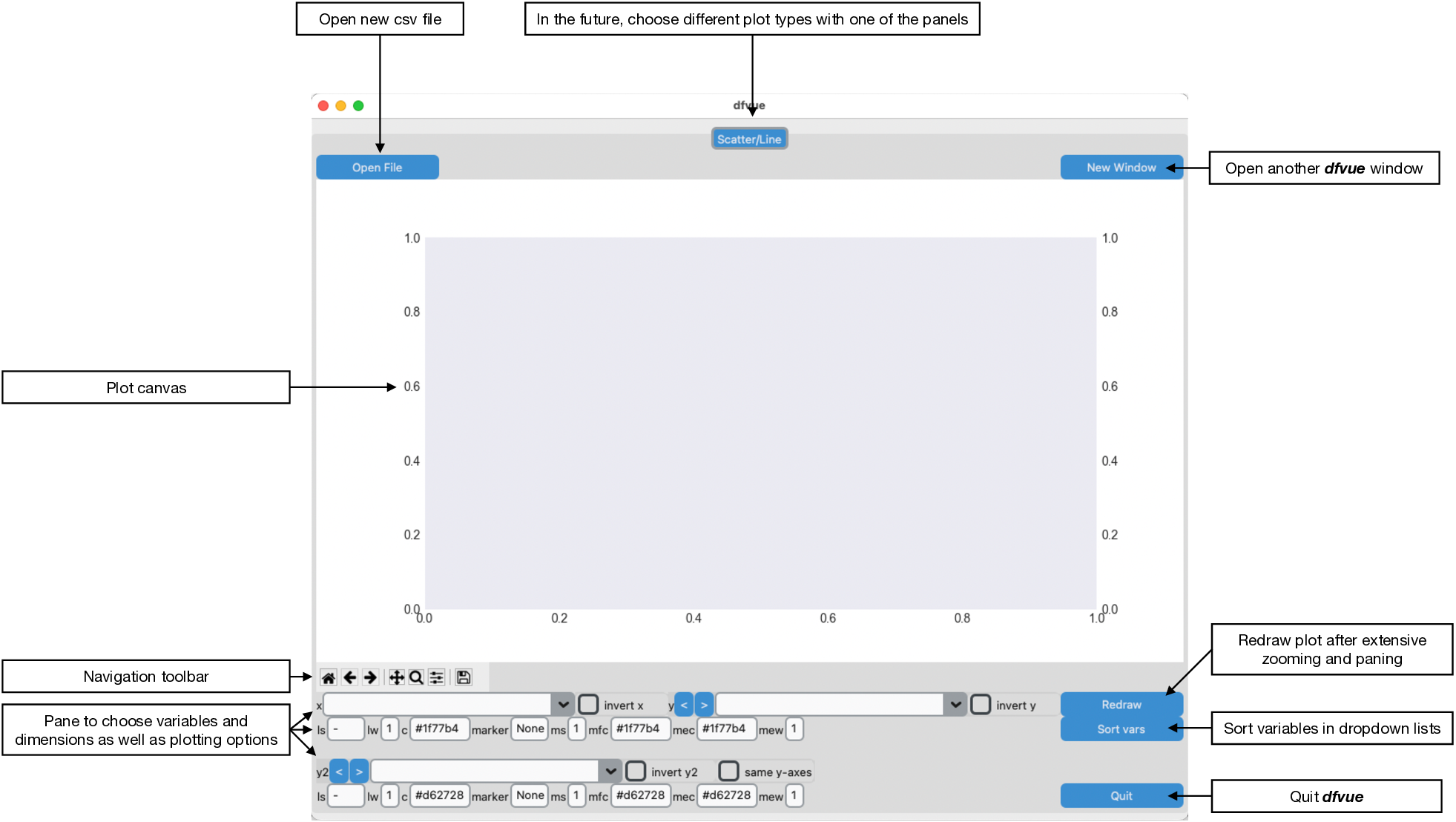
On opening, dfvue presents currently only one panel for producing
scatter/line plots. This is the look in macOS light mode (higher
resolution images can be found in the documentation):
The pane is organised in this fashion: the plotting canvas, the Matplotlib navigation toolbar and the pane, where one can choose the plotting variables and plotting options. You can open another, identical window for the same csv file with the button "New Window" on the top right. You can then also read in a new csv file in one of the windows with the button "Open File".
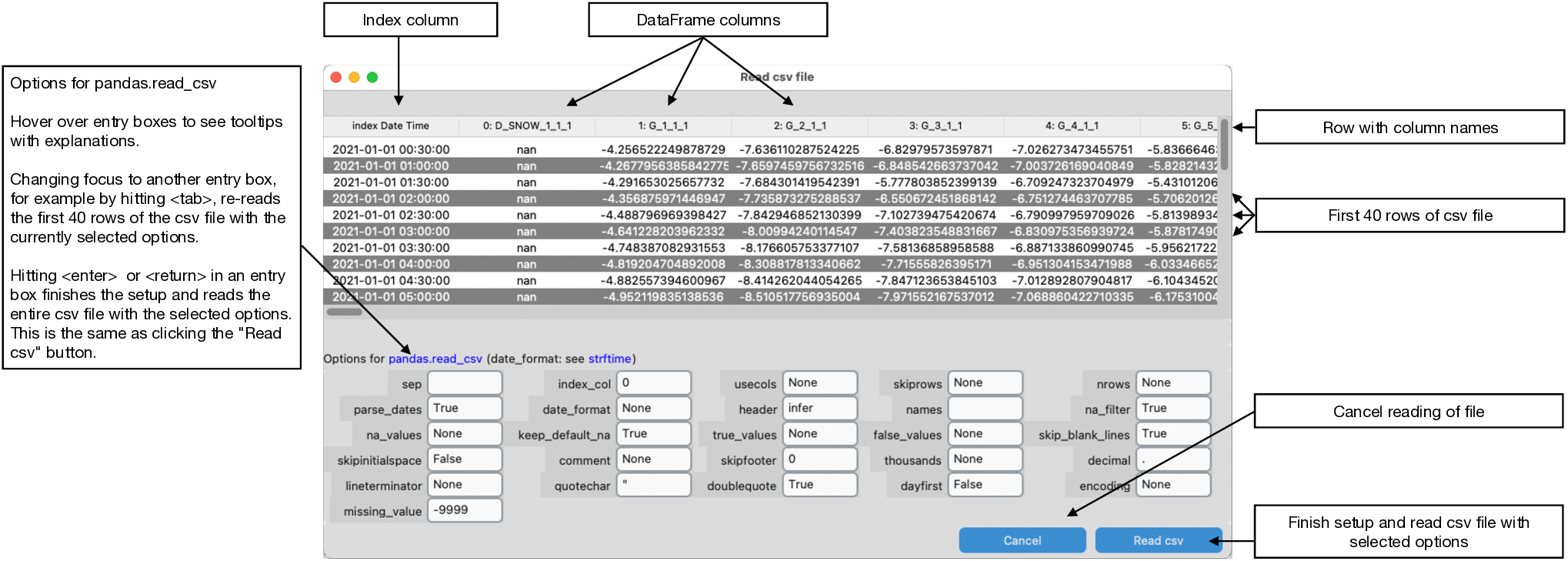
The "Read csv file" window opens when a csv file is given.
The csv file can be given on the command line:
dfvue csv_file.csvfrom within Python:
from dfvue import dfvue
dfvue('csv_file.csv')or being selected from the "Choose csv file" selector that opens when hitting the button "Open File".
The "Read csv file" window reads the first 40 rows of the csv file with pandas' read_csv method using the options given in the pane. It shows the resulting pandas.DataFrame in tabulated format. Changing focus from one option entry to another, for example by hitting the <tab> key, re-reads the first 40 rows of the csv file with pandas.read_csv using the selected options in the form. Hitting <enter> or <return> within the window reads the entire csv file using the selected options and returns to the plotting panels. This is the same than pressing the "Read csv" button in the lower right corner.
The options in the form are pandas' read_csv default options except for parse_date, which is set to True instead of False here. Hover over the entry boxes to see explanations of the options in the tooltip.
If the csv file includes a Date/Time column, it is best to set this column as the index of the pandas.DataFrame by using index_col. Correct datetime is indicated if the index has the data type datetime64[ns] in the plot panels. This is then correctly interpreted by the underlying Matplotlib when plotting, zooming, or panning the axes.
missing_value is not an option of pandas' read_csv. It is here for convenience and any number entered in missing_value will be added to pandas na_values.
The following options of pandas.read_csv can be given on the command line:
-s separator, --sep separator
Delimiter to use.
-i columns, --index_col columns
Column(s) to use as index, either given as column index
or string name.
-k rows, --skiprows rows
Line number(s) to skip (0-indexed, must include comma,
e.g. "1," for skipping the second row) or number of lines
to skip (int, without comma) at the start of the file.
-p bool/list/dict, --parse_dates bool/list/dict
boolean, if True -> try parsing the index.
list of int or names, e.g. 1,2,3
-> try parsing columns 1, 2, and 3 each as a separate
date column.
list of lists, e.g. [1,3]
-> combine columns 1 and 3 and parse as a single
date column.
dict, e.g. "foo":[1,3]
-> parse columns 1 and 3 as date and call result "foo"
-d format_string, --date_format format_string
Will parse dates according to this format.
For example: "%Y-%m-%d %H:%M%S". See
https://docs.python.org/3/library/datetime.html#strftime-and-strptime-behavior
-m missing_value, --missing_value missing_value
Missing or undefined value set to NaN. For negative values,
use long format, e.g. --missing_value=-9999.Here are some examples of csv files and the options for pandas.read_csv.
The most simple csv file would be like:
DATETIME,TA_1_1_1,RH_1_1,ALB_1_1_1 2015-01-01 00:30:00,-2.17794549084,97.2958103396,0.0 2015-01-01 01:00:00,-2.02584908489,98.2103903979,0.0
This can simply be read by setting index_col=0. The first column including date and time can simply a be a ISO8601 date, for example "2015-01-01 00:30:00" or "2015-01-01T00:30:00", or be given by date_format, which would be "%Y-%m-%d %H:%M:%S" in this case. See the documentation of pandas.to_datetime or strftime.
Command line options would be:
dfvue -i 0 csv-file
or
dfvue -i 0 -d "%Y-%m-%d %H:%M:%S" csv-file
A common practice is to put a special value for measurement errors or similar such as -9999:
DATETIME,TA_1_1_1,RH_1_1,ALB_1_1_1 2015-01-01 00:30:00,-2.17794549084,97.2958103396,-9999 2015-01-01 01:00:00,-2.02584908489,98.2103903979,-9999
This can be read by setting missing_value=-9999. On the command line, this is:
dfvue -i 0 --missing_value=-9999 csv-file
or
dfvue -i 0 -d "%Y-%m-%d %H:%M:%S" --missing_value=-9999 csv-file
You have to use the long form --missing_value=-9999 instead of the short form -m -9999 in case of negative missing values because the command line would interpret -9999 as a separate option in the second case and would fail.
Date and time information can be given in different formats, for example:
Date;rho H1 (kg/m3);alb H1 (-);T_Psy H1 (degC);WS_EC H1 (m/s);Prec H1 (mm/30min) 01.01.2015 00:30;97.2958103396;-9999;-2.17794549084 01.01.2015 01:00;98.2103903979,-9999;-2.02584908489
which can be read by setting the date format: date_format=%d.%m.%Y %H:%M, index_col=0, missing_value=-9999, as well as the field separator sep=;. On the the command line, this is:
dfvue -s ";" -i 0 -d "%d.%m.%Y %H:%M" --missing_value=-9999 csv-file
Or in FLUXNET / ICOS / europe-fluxdata.eu format with a second row that shows the variable units:
TIMESTAMP_END,TA_1_1_1,RH_1_1_1,ALB_1_1_1 YYYYMMDDhhmm,degC,%,adimensional 201501010030,-2.17794549084,97.2958103396,-9999 201501010100,-2.02584908489,98.2103903979,-9999
which is read with date_format=%Y%M%d%H%M, index_col=0, skiprows=1,, and missing_value=-9999. Note the comma after "1" in skiprows. Without the command, skiprows would be the number of rows to skip at the beginning, i.e. the first row, which would be wrong. The comma indicates that skiprows is a list and hence a list of row indexes, that means 1 here and thus skip the second row. This would be on the command line
dfvue -i 0 -d "%Y%m%d%H%M" --skiprows=1, --missing_value=-9999 csv-file
Date and time information can also be in different columns. Here the second column is the day-of-the-year:
year,jday,hour,min,tair,rhair,albedo 2015,1,0,30,-2.17794549084,97.2958103396,-9999 2015,1,1,0,-2.02584908489,98.2103903979,-9999
which can be read by setting parse_dates=[0,1,2,3], index_col=0, and date_format=%Y %j %H %M, as well as missing_value=-9999. Note the brackets "[]" around parse_dates. Without brackets it would parse columns 0, 1, 2, and 3 each as a separate date column, whereas with brackets it combines columns 0, 1, 2, and 3 and parses it as a single date column, with index "0". It will use a space between column entries. Hence index_col=0 sets this combined column as the index, parsing the dates with the format "%Y %j %H %M" with spaces between the strftime formats.
On the command line, this would be:
dfvue -i 0 -p [0,1,2,3] -d "%Y %j %H %M" --missing_value=-9999 csv-file
If you want to have spaces in the list of parse_dates on the command line, you have to use the long form: --parse_dates="[0, 1, 2, 3]".
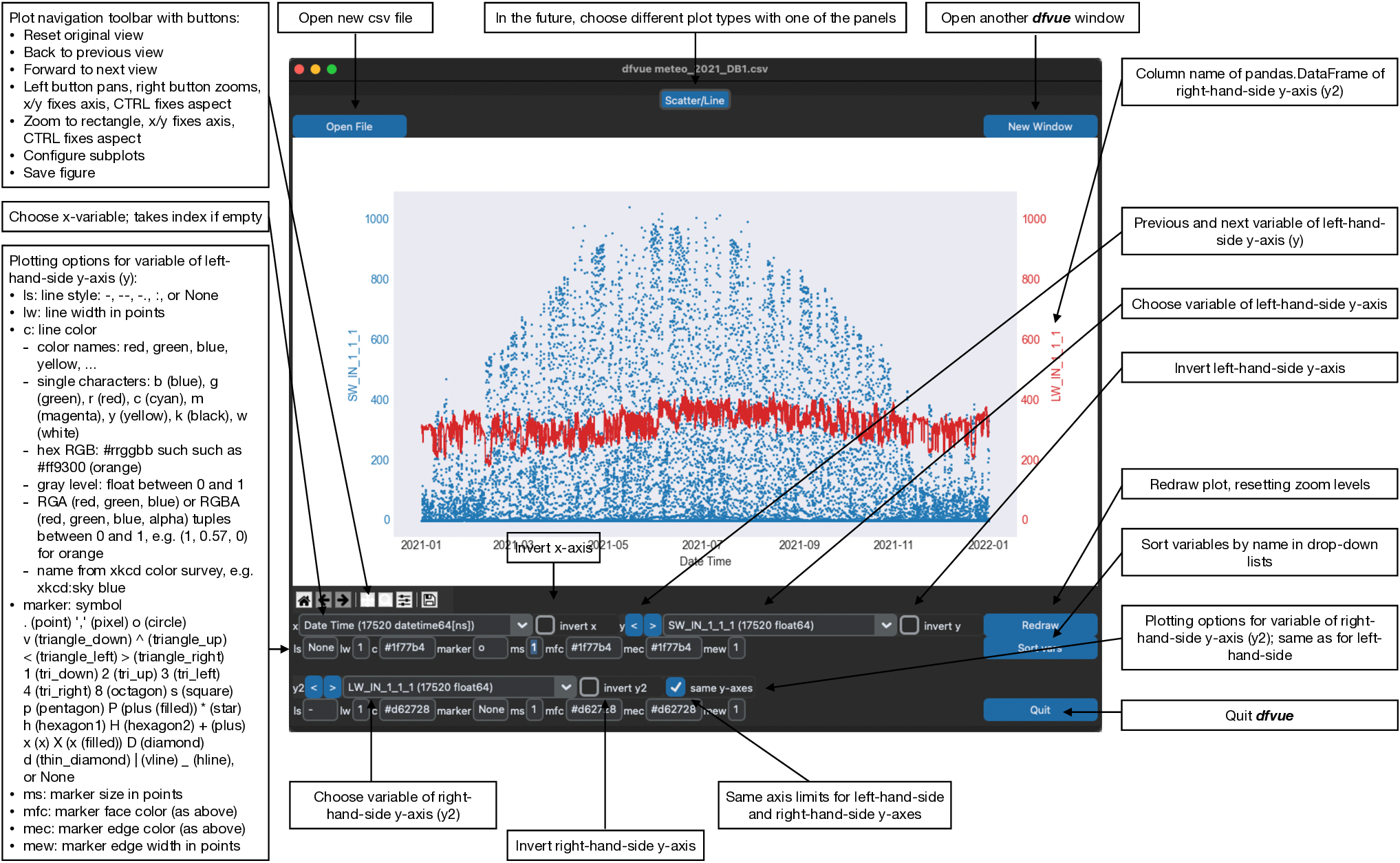
Here is the Scatter/Line panel in macOS dark mode, describing all buttons, sliders, entry boxes, spinboxes, and menus:
The default plot is a line plot with solid lines (line style 'ls' is '-'). One can set line style 'ls' to None and set a marker symbol, e.g. 'o' for circles, to get a scatter plot. A large variety of line styles, marker symbols and color notations are supported.
dfvue is an application written in Python. It can be installed
with pip:
python -m pip install dfvueor via Conda:
conda install -c conda-forge dfvuedfvue uses CustomTkinter if it is installed. CustomTkinter is not on Conda. One might install dfvue with pip in a conda environment to use CustomTkinter:
conda install pip
python -m pip install dfvueIf this looks ugly on Linux (see this thread), pip uninstall customtkinter, or pip uninstall dfvue and reinstall it with conda, which then uses the Azure theme by rdbende.
If the fonts in dfvue (and any other tkinter GUI) still look
ugly, one can try to reinstall Tk with FreeType support via Xft:
conda install -c conda-forge tk=*=xft_*dfvue is distributed under the MIT License. See the LICENSE file
for details.
Copyright (c) 2023- Matthias Cuntz
dfvue uses CustomTkinter by Tom Schimansky if installed, and
otherwise the Azure theme (v2.0) by rdbende, for example in conda
environments.