Efficient and comprehensive metadata acquisition from NCBI databases (includes SRA).
NCBImeta is a command-line application that retrieves and organizes metadata from the National Centre for Biotechnology Information (NCBI). While the NCBI web browser experience allows filtered searches, the output does not facilitate inter-record comparison or bulk record retrieval. NCBImeta tackles this issue by creating a local database of NCBI metadata constructed by user-defined search criteria and customizable metadata columns. The output of NCBImeta, optionally a SQLite database or text files, can then be used by computational biologists for applications such as record filtering, project discovery, sample interpretation, or meta-analyses of published work.
NCBImeta is written in Python 3 and supported on Linux and macOS. Dependencies that will be installed are listed in requirements.txt.
Check all Python versions and OS with verified build status
There are three installation options for NCBImeta:
conda install -c bioconda ncbimeta
pip install NCBImeta
If you have both Python2 and Python3 installed, using pip3 may be beneficial instead:
pip3 install NCBImeta
git clone https://github.com/ktmeaton/NCBImeta.git
cd NCBImeta
python setup.py install
Test that the installation was successful:
NCBImeta.py --version
Download the NCBImeta github repository to get access to the example configuration files:
git clone https://github.com/ktmeaton/NCBImeta.git
cd NCBImeta
Download a selection of genomic metadata pertaining to the plague pathogen Yersinia pestis.
NCBImeta.py --flat --config example/config.yaml
(Note: The 'quick' start config file forces slow downloads to accommodate users with slow internet. For faster record retrieval, please see the Config File README to start editing config files.)
Example output of the command-line interface (v0.6.1):
NCBImetaAnnotateReplace.py --database example/yersinia_pestis_db.sqlite --annotfile example/annot.txt --table BioSample
Note that the first column of your annotation file MUST be a column that is unique to each record. An Accession number or ID is highly recommended. The column headers in your annotation file must also exactly match the names of your columns in the database.
NCBImetaAnnotateReplace.py, as the name implies, replaces the existing annotation with the data in your custom metadata file. Alternatively, the script NCBImetaAnnotateConcatenate.py will concatenate your custom metadata with the pre-existing value in the database cell (separated by a semi-colon).
NCBImetaAnnotateConcatenate.py --database example/yersinia_pestis_db.sqlite --annotfile example/annot.txt --table BioSample
NCBImetaJoin.py --database example/yersinia_pestis_db.sqlite --final Master --anchor BioSample --accessory "BioProject Assembly SRA Nucleotide" --unique "BioSampleAccession BioSampleAccessionSecondary BioSampleBioProjectAccession"
The rows of the output "Master" table will be from the anchor table "BioSample", with additional columns added in from the accessory tables "BioProject", "Assembly", "SRA", and "Nucleotide". Unique accession numbers for BioSample (both primary and secondary) and BioProject allow this join to be unambiguous.
NCBImetaExport.py --database example/yersinia_pestis_db.sqlite --outputdir example/
Each table within the database will be exported to its own tab-separated .txt file in the specified output directory.
- Explore your database text files using a spreadsheet viewer (Microsoft Excel, Google Sheets, etc.)
- Browse your SQLite database using DB Browser for SQLite (https://sqlitebrowser.org/)
- Use the columns with FTP links to download your data files of interest.
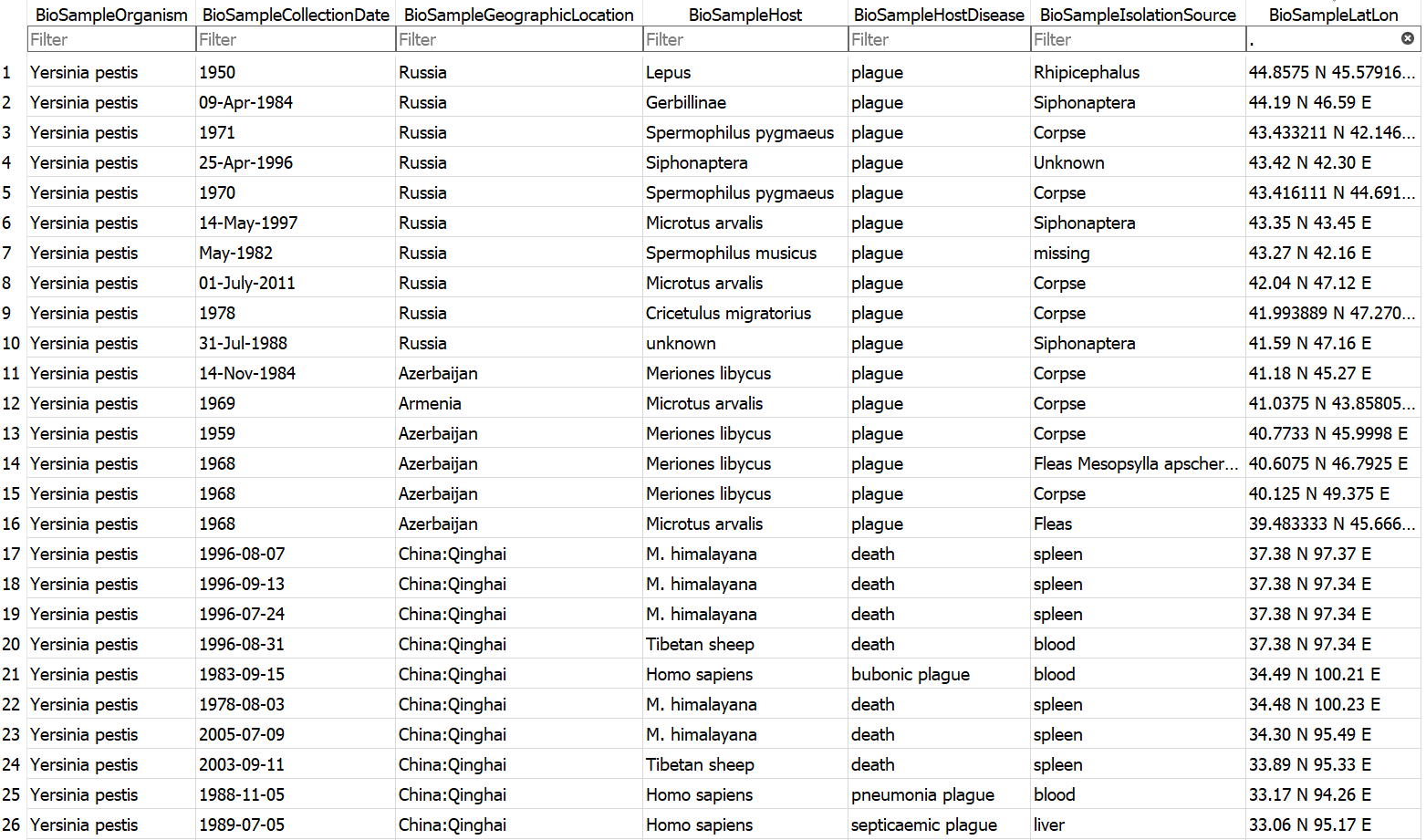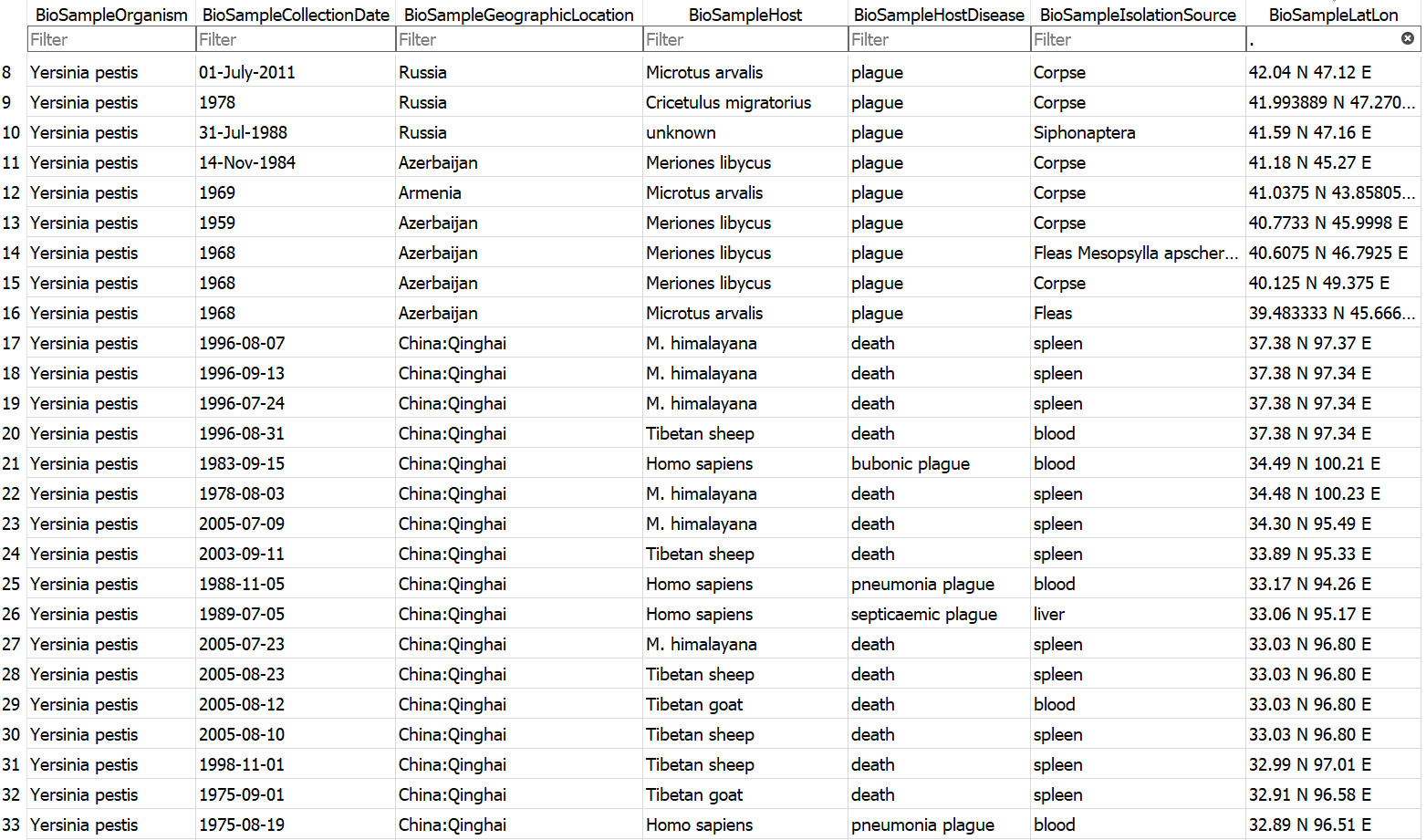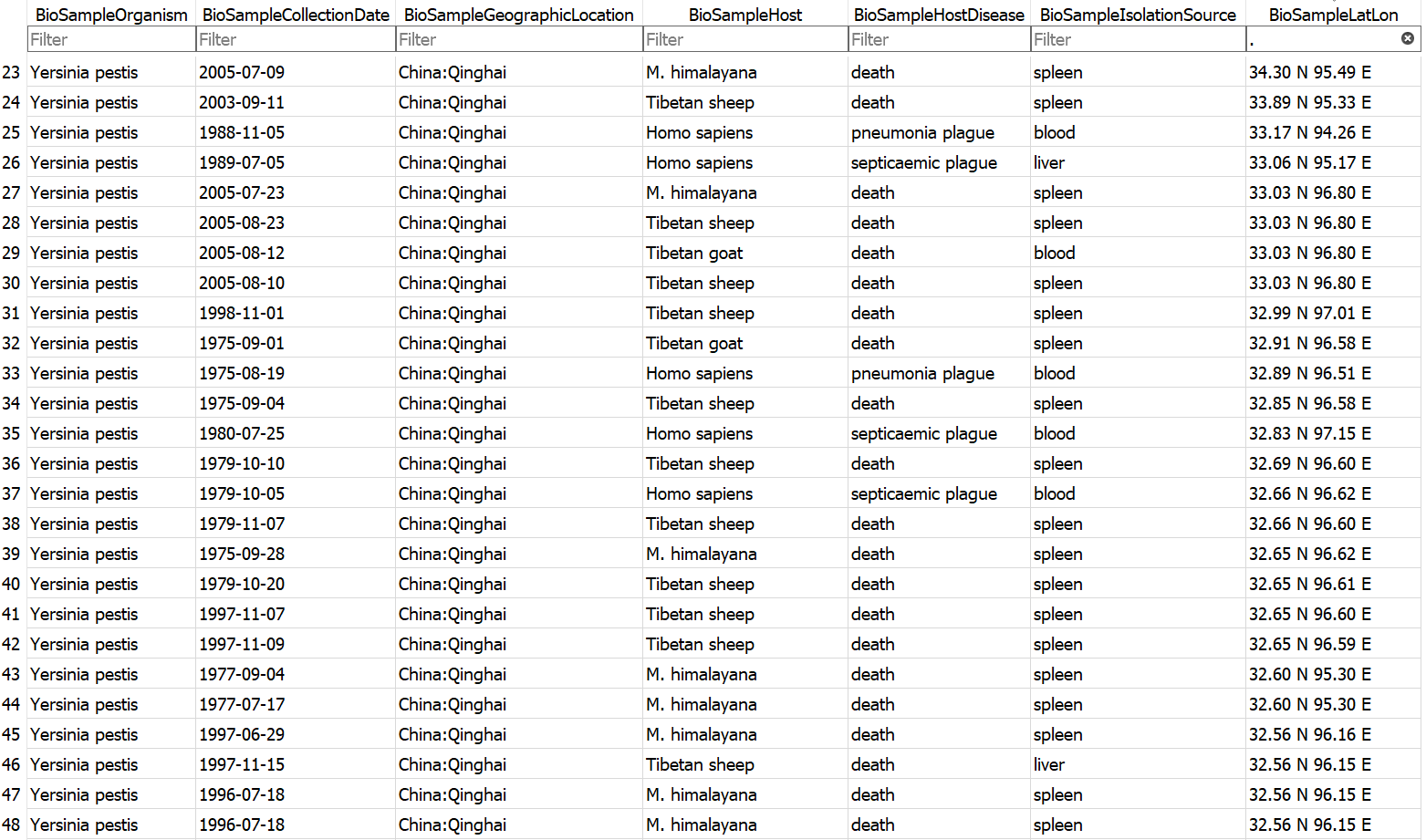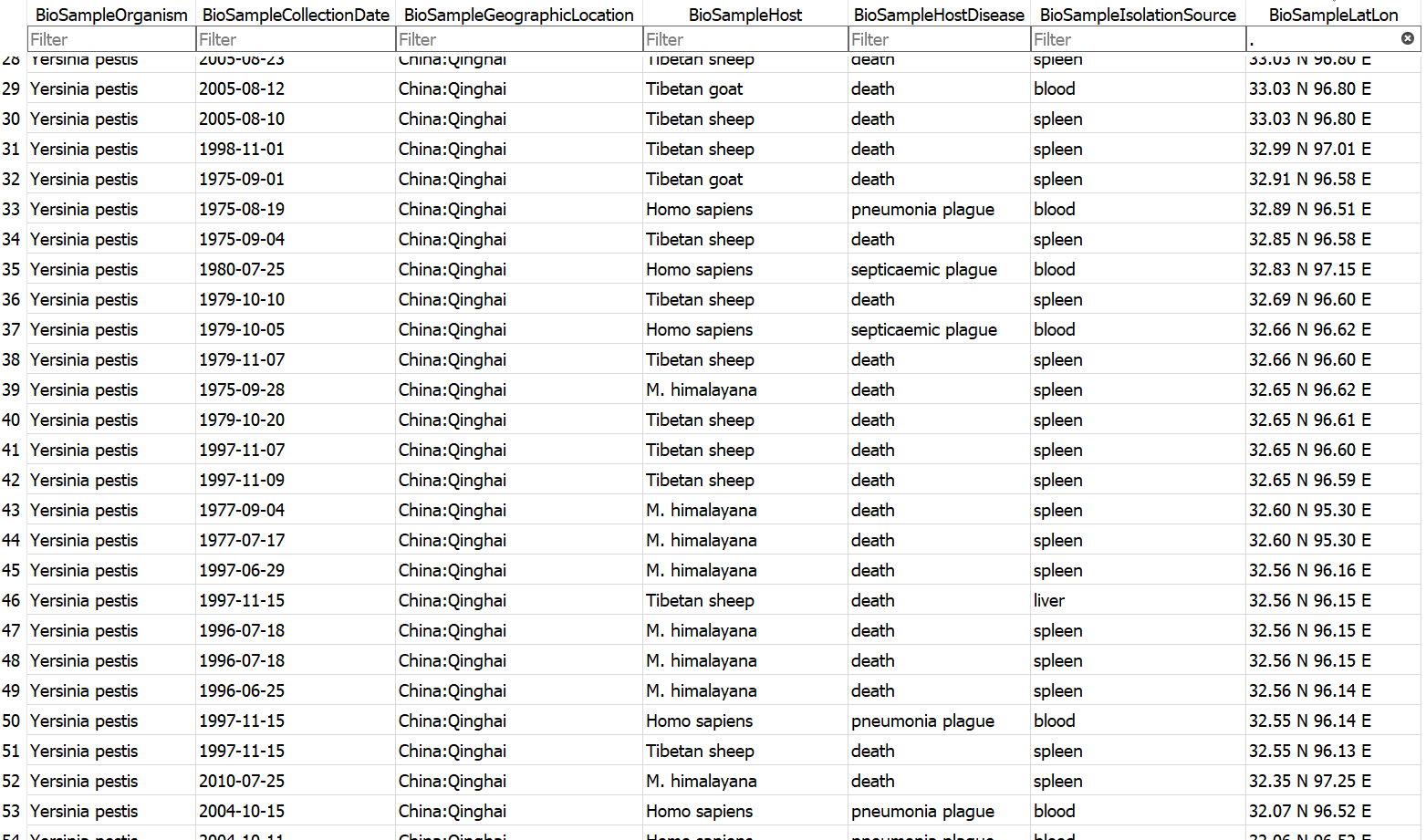
Example database output (a subset of the BioSample table)
- Assembly
- BioProject
- BioSample
- Nucleotide
- SRA
- Pubmed
- Project v0.6.4 - "Bobby Tables": SQL Security Update
- Project v0.6.5 - "NextFlow": Improved NextFlow integration and quiet mode
- Project v0.6.x - "Read the Docs": Full documentation pages on Read the Docs
To get started with customizing the search terms, database, and metadata fields, please read:
Please submit your questions, suggestions, and bug reports to the Issue Tracker.
Please do not hesitate to post any manner of curiosity in the "Issues" tracker :) User-feedback and ideas are the most valuable resource for emerging software.
- Fork it!
- Create your feature branch:
git checkout -b my-new-feature - Commit your changes:
git commit -am 'Add some feature' - Push to the branch:
git push origin my-new-feature - Submit a pull request!
Eaton, K. (2020). NCBImeta: efficient and comprehensive metadata retrieval from NCBI databases. Journal of Open Source Software, 5(46), 1990, https://doi.org/10.21105/joss.01990
Author: Katherine Eaton ([email protected])