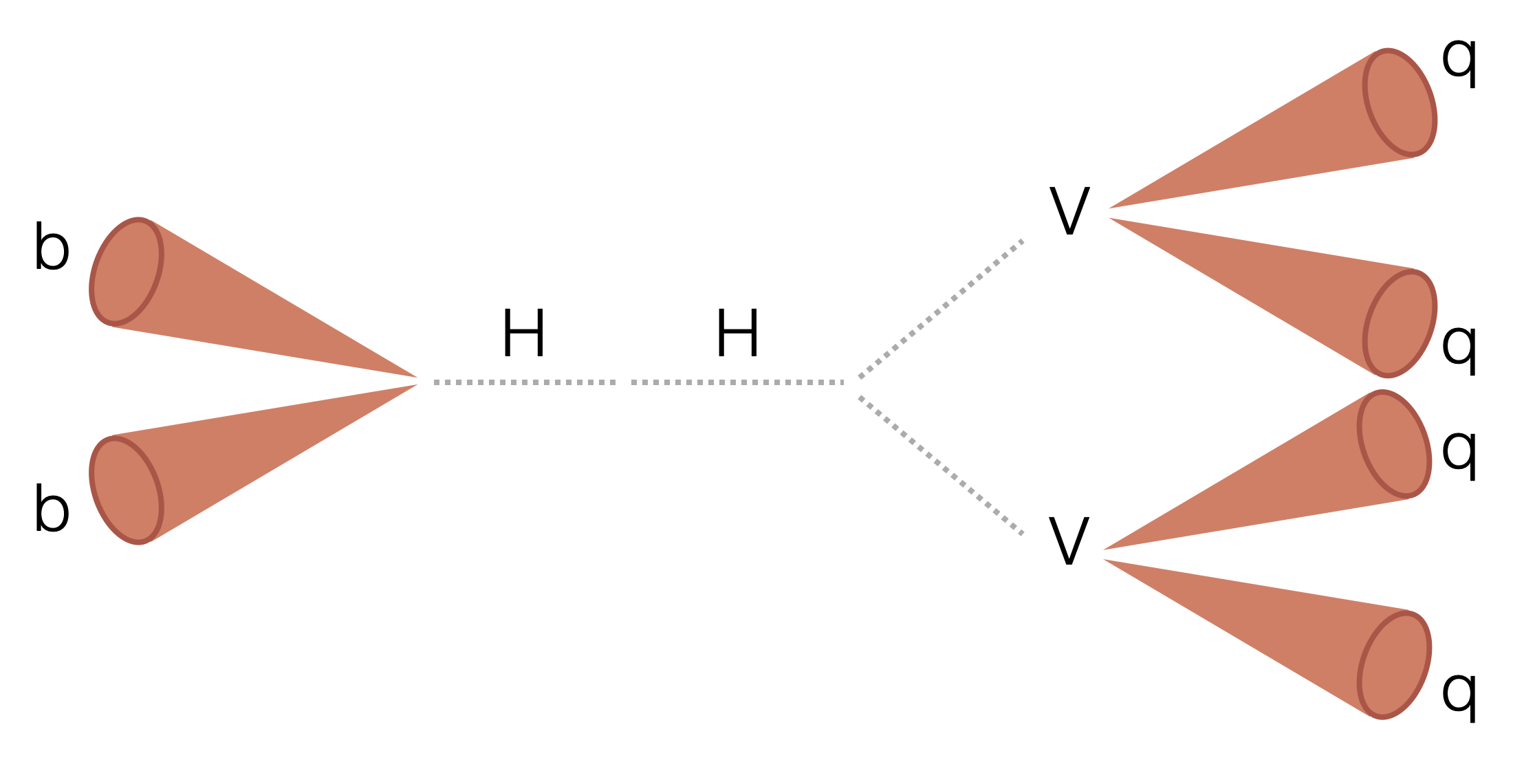
Search for two boosted (high transverse momentum) Higgs bosons (H) decaying to two beauty quarks (b) and two vector bosons (V). The majority of the analysis uses a columnar framework to process input tree-based NanoAOD files using the coffea and scikit-hep Python libraries.
General note: Coffea-casa is faster and more convenient, however still somewhat experimental so for large of inputs and/or processors which may require heavier cpu/memory usage (e.g. bbVVSkimmer) condor is recommended.
For coffea-casa:
- after following instructions ^ set up an account, open the coffea-casa GUI (https://cmsaf-jh.unl.edu) and create an image
- open
src/runCoffeaCasa.ipynb - import your desired processor, specify it in the
run_uproot_jobfunction, and specify your filelist - run the first three cells
git clone https://github.com/rkansal47/HHbbVV/
cd HHbbVV
TAG=Aug18_skimmer
python src/condor/submit.py --processor skimmer --tag $TAG --files-per-job 20 # will need python3 (recommended to set up via miniconda)
for i in condor/$TAG/*.jdl; do condor_submit $i; doneTo test locally first (recommended), can do e.g.:
mkdir outfiles
python -W ignore src/run.py --starti 0 --endi 1 --year 2017 --processor skimmer --executor iterative --samples HWW --subsamples GluGluToHHTobbVV_node_cHHH1_pn4qApplies pre-selection cuts, runs inference with our new HVV tagger, and saves unbinned branches as parquet files.
Parquet and pickle files will be saved in the eos directory of specified user at path ~/eos/bbVV/skimmer/<tag>/<sample_name>/<parquet or pickles>. Pickles are in the format {'nevents': int, 'cutflow': Dict[str, int]}.
To test locally:
python -W ignore src/run.py --processor skimmer --samples HWW --subsamples GluGluToHHTobbVV_node_cHHH1_pn4q --save-ak15 --starti 0 --endi 1
Jobs
python src/condor/submit_from_yaml.py --tag Jul24 --processor skimmer --save-ak15 --submit --yaml src/condor/submit_configs/skimmer_inputs_07_24.yaml
# Submit (if not use --submit flag)
nohup bash -c 'for i in condor/'"${TAG}"'/*.jdl; do condor_submit $i; done' &> tmp/submitout.txt &Applies a loose pre-selection cut, saves ntuples with training inputs.
To test locally (in singularity):
python -W ignore src/run.py --year 2017 --starti 300 --endi 301 --samples HWWPrivate --subsamples jhu_HHbbWW --processor input --label AK15_H_VV
python -W ignore src/run.py --year 2017 --starti 300 --endi 301 --samples QCD --subsamples QCD_Pt_1000to1400 --processor input --label AK15_QCD --njets 1 --maxchunks 1Jobs:
python src/condor/submit_from_yaml.py --tag Jul21 --processor input --save-ak15 --yaml src/condor/submit_configs/tagger_inputs_07_21.yaml
To submit add --submit flag.
Command for copying directories to PRP in background (krsync)
(you may need to install rsync in the PRP pod first if it's been restarted)
cd ~/eos/bbVV/input/${TAG}_${JETS}/2017
mkdir ../copy_logs
for i in *; do echo $i && sleep 3 && (nohup sh -c "krsync -av --progress --stats $i/root/ $HWWTAGGERDEP_POD:/hwwtaggervol/training/$FOLDER/$i" &> ../copy_logs/$i.txt &) done```