Main features • Installation • Overview • Databases • Data model • Example workflow • Analysis across trials • Tests • Acknowledgements • Future
The package ctrdata provides functions for retrieving (downloading)
information on clinical trials from public registers, and for
aggregating and analysing this information; it can be used for the
- EU Clinical Trials Register (“EUCTR”, https://www.clinicaltrialsregister.eu/)
- EU Clinical Trials Information System (“CTIS”, https://euclinicaltrials.eu/, see example)
- ClinicalTrials.gov (“CTGOV2”, see example)
- ISRCTN Registry (https://www.isrctn.com/)
The motivation is to investigate and understand trends in design and
conduct of trials, their availability for patients and to facilitate
using their detailed results for research and meta-analyses. ctrdata
is a package for the R system, but other
systems and tools can be used with the databases created with the
package. This README was reviewed on 2024-08-08 for version 1.19.1.9000
(recent improvements: removed external dependencies; refactored
dbGetFieldsIntoDf();
🔔 retrieve historic CTGOV2 versions; adapt to relaunched CTIS).
- Protocol- and results-related trial information is easily downloaded:
Users define a query in a register’s web interface, then copy the URL
and enter it into
ctrdatawhich retrieves in one go all trials found. A script can automate copying the query URL from all registers. Personal annotations can be made when downloading trials. Also, trial documents and historic versions available in registers on trials can be downloaded. - Downloaded trial information is transformed and stored in a collection
of a document-centric database, for fast and offline access.
Information from different registers can be accumulated in a single
collection. Uses
DuckDB,PostgreSQL,RSQLiteorMongoDB, via R packagenodbi: see section Databases below. Easily re-run any previous query in a collection to retrieve and update trial records. - For analyses, convenience functions in
ctrdataallow find synonyms of an active substance, to identify unique (de-duplicated) trial records across all registers, to merge and recode fields as well as to easily access deeply-nested fields. Analysis can be done withR(see vignette) or other systems, using theJSON-structured information in the database.
Remember to respect the registers’ terms and conditions (see
ctrOpenSearchPagesInBrowser(copyright = TRUE)). Please cite this
package in any publication as follows: “Ralf Herold (2024). ctrdata:
Retrieve and Analyze Clinical Trials in Public Registers. R package
version 1.19.1, https://cran.r-project.org/package=ctrdata”.
Package ctrdata has been used for unpublished work and for:
- Lasch et al. (2022) The Impact of COVID‐19 on the Initiation of Clinical Trials in Europe and the United States. Clinical Pharmacology & Therapeutics https://doi.org/10.1002/cpt.2534
- Sood et al. Managing the Evidence Infodemic: Automation Approaches Used for Developing NICE COVID-19 Living Guidelines. medRxiv https://doi.org/10.1101/2022.06.13.22276242 🔔
- Blogging on Innovation coming to paediatric research
- Cancer Research UK (2017) The impact of collaboration: The value of UK medical research to EU science and health
Package ctrdata is on
CRAN and on
GitHub. Within
R, use the following commands to install
package ctrdata:
# Install CRAN version:
install.packages("ctrdata")
# Alternatively, install development version:
install.packages("devtools")
devtools::install_github("rfhb/ctrdata", build_vignettes = TRUE)These commands also install the package’s dependencies (jsonlite,
httr, curl, clipr, xml2, nodbi, stringi, tibble,
lubridate, jqr, dplyr, zip and V8).
This is optional; it works with all registers supported by ctrdata but
is recommended for CTIS because the URL in the web browser does not
reflect the parameters the user specified for querying this register.
In the web browser, install the Tampermonkey browser
extension, click on “New user script”
and then on “Tools”, then enter into “Import from URL” this URL:
https://raw.githubusercontent.com/rfhb/ctrdata/master/tools/ctrdataURLcopier.js
and last click on “Install”.
The browser extension can be disabled and enabled by the user. When
enabled, the URLs to all user’s queries in the registers are
automatically copied to the clipboard and can be pasted into the
queryterm = ... parameter of function
ctrLoadQueryIntoDb()
The functions are listed in the approximate order of use in a user’s workflow (in bold, main functions). See also the package documentation overview.
| Function name | Function purpose |
|---|---|
ctrOpenSearchPagesInBrowser() |
Open search pages of registers or execute search in web browser |
ctrFindActiveSubstanceSynonyms() |
Find synonyms and alternative names for an active substance |
ctrGetQueryUrl() |
Import from clipboard the URL of a search in one of the registers |
ctrLoadQueryIntoDb() |
Retrieve (download) or update, and annotate, information on trials from a register and store in a collection in a database |
dbQueryHistory() |
Show the history of queries that were downloaded into the collection |
dbFindIdsUniqueTrials() |
Get the identifiers of de-duplicated trials in the collection |
dbFindFields() |
Find names of variables (fields) in the collection |
dbGetFieldsIntoDf() |
Create a data frame (or tibble) from trial records in the database with the specified fields |
dfTrials2Long() |
Transform the data.frame from dbGetFieldsIntoDf() into a long name-value data.frame, including deeply nested fields |
dfName2Value() |
From a long name-value data.frame, extract values for variables (fields) of interest (e.g., endpoints) |
dfMergeVariablesRelevel() |
Merge variables into a new variable, optionally map values to a new set of levels |
Package ctrdata retrieves trial information and stores it in a
database collection, which has to be given as a connection object to
parameter con for several ctrdata functions; this connection object
is created in slightly different ways for the four supported database
backends that can be used with ctrdata as shown in the table. For a
speed comparison, see the nodbi
documentation.
Besides ctrdata functions below, any such a connection object can
equally be used with functions of other packages, such as nodbi (last
row in table) or, in case of MongoDB as database backend, mongolite
(see vignettes).
| Purpose | Function call |
|---|---|
| Create SQLite database connection | dbc <- nodbi::src_sqlite(dbname = "name_of_my_database", collection = "name_of_my_collection") |
| Create MongoDB database connection | dbc <- nodbi::src_mongo(db = "name_of_my_database", collection = "name_of_my_collection") |
| Create PostgreSQL database connection | dbc <- nodbi::src_postgres(dbname = "name_of_my_database"); dbc[["collection"]] <- "name_of_my_collection" |
| Create DuckDB database connection | dbc <- nodbi::src_duckdb(dbdir = "name_of_my_database", collection = "name_of_my_collection") |
Use connection with ctrdata functions |
ctrdata::{ctrLoadQueryIntoDb, dbQueryHistory, dbFindIdsUniqueTrials, dbFindFields, dbGetFieldsIntoDf}(con = dbc, ...) |
Use connection with nodbi functions |
e.g., nodbi::docdb_query(src = dbc, key = dbc$collection, ...) |
Package ctrdata uses the data models that are implicit in data
retrieved from the different registers. No mapping is provided for any
register’s data model to a putative target data model. The reasons
include that registers’ data models are notably evolving over time and
that there are only few data fields with similar values and meaning
between the registers.
Thus, the handling of data from different models of registers is to be done at the time of analysis. This approach allows a high level of flexibility, transparency and reproducibility. See examples in the help text for function dfMergeVariablesRelevel() and section Analysis across trials below for how to align related fields from different registers for a joint analysis.
In any of the NoSQL
databases,
one clinical trial is one document, corresponding to one row in a
SQLite, PostgreSQL or DuckDB table, and to one document in a
MongoDB collection. The NoSQL backends allow documents to have
different structures, which is used here to accommodate the different
data models of registers. Package ctrdata stores in every such
document:
- field
_idwith the trial identification as provided by the register from which it was retrieved - field
ctrnamewith the name of the register (EUCTR,CTGOV,CTGOV2,ISRCTN,CTIS) from which that trial was retrieved - field
record_last_importwith the date and time when that document was last updated usingctrLoadQueryIntoDb() - only for
CTGOV2: objecthistorywith a historic version of the trial and withhistory_version, which contains the fieldsversion_number(starting from 1) andversion_date - all original fields as provided by the register for that trial (see examples below)
For visualising the data structure for a trial, see this vignette section.
- Install R package ctrdata
- Retrieve clinical trial information
- Summarise and analyse clinical trial information
The aim is to download protocol-related trial information and tabulate the trials’ status of conduct.
- Attach package
ctrdata:
library(ctrdata)- See help to get started with
ctrdata:
help("ctrdata")- Information on trial registers and how they can be used with
ctrdata(last updated 2024-06-23):
help("ctrdata-registers")- Open registers’ advanced search pages in browser:
ctrOpenSearchPagesInBrowser()
# Please review and respect register copyrights:
ctrOpenSearchPagesInBrowser(copyright = TRUE)-
Adjust search parameters and execute search in browser
-
When trials of interest are listed in browser, copy the address from the browser’s address bar to the clipboard (you can automate this, see here)
-
Search used in this example: https://www.clinicaltrialsregister.eu/ctr-search/search?query=cancer&age=under-18&phase=phase-one&status=completed
-
Get address from clipboard:
q <- ctrGetQueryUrl()
# * Using clipboard content as register query URL:
# https://www.clinicaltrialsregister.eu/ctr-search/search?query=cancer&
# age=under-18&phase=phase-one&status=completed
# * Found search query from EUCTR: query=cancer&age=under-18&phase=phase-one&status=completed
q
# query-term query-register
# 1 query=cancer&age=under-18&phase=phase-one&status=completed EUCTR🔔 Queries in the trial registers can automatically copied to the clipboard (including for “CTIS”, where the URL does not show the query) using our solution here.
- Retrieve protocol-related information, transform and save to database:
The database collection is specified first, using nodbi (see above for
how to specify PostgreSQL, RSQlite, DuckDB or MongoDB as
backend, see section
Databases).
Then, trial information is retrieved and loaded into the collection:
# Connect to (or create) an SQLite database
# stored in a file on the local system:
db <- nodbi::src_sqlite(
dbname = "some_database_name.sqlite_file",
collection = "some_collection_name"
)
# Retrieve trials from public register:
ctrLoadQueryIntoDb(
queryterm = q,
euctrresults = TRUE,
con = db
)
# * Found search query from EUCTR: query=cancer&age=under-18&phase=phase-one&status=completed
# * Checking trials in EUCTR...
# Retrieved overview, multiple records of 104 trial(s) from 6 page(s) to be downloaded (estimate: 10 MB)
# (1/3) Downloading trials...
# Note: register server cannot compress data, transfer takes longer (estimate: 100 s)
# Download status: 6 done; 0 in progress. Total size: 8.91 Mb (100%)... done!
# (2/3) Converting to NDJSON (estimate: 2 s)...
# (3/3) Importing records into database...
# = Imported or updated 418 records on 104 trial(s)
# * Checking results if available from EUCTR for 104 trials:
# (1/4) Downloading results...
# Download status: 104 done; 0 in progress. Total size: 59.72 Mb (100%)... done!
# Download status: 28 done; 0 in progress. Total size: 112.71 Kb (100%)... done!
# - extracting results (. = data, F = file[s] and data, x = none):
# F . F F . F . . F . . . F F . . . . . . . . . . . . . . . . . . F . . . . . . F . .
# . F . . . . . . . . . F . . . . F . . . . . F . . . . . . . . . . .
# (2/4) Converting to NDJSON (estimate: 8 s)...
# (3/4) Importing results into database (may take some time)...
# (4/4) Results history: not retrieved (euctrresultshistory = FALSE)
# = Imported or updated results for 76 trials
# No history found in expected format.
# Updated history ("meta-info" in "some_collection_name")
# $n
# [1] 418Under the hood, EUCTR plain text and XML files from EUCTR, CTGOV, ISRCTN
are converted using Javascript via V8 in R into NDJSON, which is
imported into the database collection.
- Analyse
Tabulate the status of trials that are part of an agreed paediatric
development program (paediatric investigation plan, PIP). ctrdata
functions return a data.frame (or a tibble, if package tibble is
loaded).
# Get all records that have values in the fields of interest:
result <- dbGetFieldsIntoDf(
fields = c(
"a7_trial_is_part_of_a_paediatric_investigation_plan",
"p_end_of_trial_status",
"a2_eudract_number"
),
con = db
)
# Find unique (deduplicated) trial identifiers for trials that have more than
# one record, for example for several EU Member States or in several registers:
uniqueids <- dbFindIdsUniqueTrials(con = db)
# Searching for duplicate trials...
# - Getting all trial identifiers (may take some time), 418 found in collection
# - Finding duplicates among registers' and sponsor ids...
# - 314 EUCTR _id were not preferred EU Member State record for 104 trials
# - Keeping 104 / 0 / 0 / 0 / 0 records from EUCTR / CTGOV / CTGOV2 / ISRCTN / CTIS
# = Returning keys (_id) of 104 records in collection "some_collection_name"
# Keep only unique / de-duplicated records:
result <- subset(
result,
subset = `_id` %in% uniqueids
)
# Tabulate the selected clinical trial information:
with(
result,
table(
p_end_of_trial_status,
a7_trial_is_part_of_a_paediatric_investigation_plan
)
)
# a7_trial_is_part_of_a_paediatric_investigation_plan
# p_end_of_trial_status FALSE TRUE
# Completed 50 23
# GB - no longer in EU/EEA 1 1
# Ongoing 4 3
# Prematurely Ended 2 3
# Restarted 0 1
# Temporarily Halted 1 1
# Trial now transitioned 1 0- Add records from another register (CTGOV2) into the same collection
The new website and API introduced in July 2023
(https://www.clinicaltrials.gov/) is supported by ctrdata since
mid-2023 and identified in ctrdata as CTGOV2.
On 2024-06-25, CTGOV has retired the classic website and API used by
ctrdata since 2015. To support users, ctrdata however automatically
translates and redirects queries to the current website. This helps with
automatically updating previously loaded queries
(ctrLoadQueryIntoDb(querytoupdate = <n>)), manually migrating queries
and reproducible work on clinical trials information. Going forward,
users are recommended to change to use CTGOV2 queries.
As regards study data, important differences exist between field names
and contents of information retrieved using CTGOV or CTGOV2; see the
schema for study protocols in
CTGOV,
the schema for study
results
and the Study Data Structure for
CTGOV2.
For more details, call help("ctrdata-registers"). This is one of the
reasons why ctrdata handles the situation as if these were two
different registers and will continue to identify the current API as
register = "CTGOV2", to support the analysis stage.
Note that loading trials with ctrdata overwrites the previous record
with CTGOV2 data, whether the previous record was retrieved using
CTGOV or CTGOV queries.
- Search used in this example: https://www.clinicaltrials.gov/search?cond=Neuroblastoma&aggFilters=ages:child,results:with,studyType:int
# Retrieve trials from another register:
ctrLoadQueryIntoDb(
queryterm = "cond=Neuroblastoma&aggFilters=ages:child,results:with,studyType:int",
register = "CTGOV2",
con = db
)
# * Appears specific for CTGOV REST API 2.0
# * Found search query from CTGOV2: cond=Neuroblastoma&aggFilters=ages:child,results:with,studyType:int
# * Checking trials using CTGOV REST API 2.0, found 100 trials
# (1/3) Downloading in 1 batch(es) (max. 1000 trials each; estimate: 10 MB total)
# Download status: 1 done; 0 in progress. Total size: 9.19 Mb (805%)... done!
# (2/3) Converting to NDJSON...
# (3/3) Importing records into database...
# JSON file #: 1 / 1
# = Imported or updated 100 trial(s)
# Updated history ("meta-info" in "some_collection_name")
# $n
# [1] 100- Using an example from classic CTGOV:
# Retrieve trials:
ctrLoadQueryIntoDb(
queryterm = paste0(
"https://classic.clinicaltrials.gov/ct2/results?",
"cond=neuroblastoma&rslt=With&recrs=e&age=0&intr=Drug"),
con = db
)
# * Appears specific for CTGOV Classic website
# Since 2024-06-25, the classic CTGOV servers are no longer available. Package
# ctrdata has translated the classic CTGOV query URL from this call of function
# ctrLoadQueryIntoDb(queryterm = ...) into a query URL that works with the current
# CTGOV2. This is printed below and is also part of the return value of this function,
# ctrLoadQueryIntoDb(...)$url. This URL can be used with ctrdata functions. Note that
# the fields and data schema of trials differ between CTGOV and CTGOV2.
#
# Replace this URL:
#
# https://classic.clinicaltrials.gov/ct2/results?cond=neuroblastoma&rslt=With&recrs=e&age=0&intr=Drug
#
# with this URL:
#
# https://clinicaltrials.gov/search?cond=neuroblastoma&intr=Drug&aggFilters=ages:child,results:with,status:com
#
# * Found search query from CTGOV2: cond=neuroblastoma&intr=Drug&aggFilters=ages:child,results:with,status:com
# * Checking trials using CTGOV REST API 2.0, found 62 trials
# (1/3) Downloading in 1 batch(es) (max. 1000 trials each; estimate: 6.2 MB total)
# Download status: 1 done; 0 in progress. Total size: 7.12 Mb (937%)... done!
# (2/3) Converting to NDJSON...
# (3/3) Importing records into database...
# JSON file #: 1 / 1
# = Imported or updated 62 trial(s)
# Updated history ("meta-info" in "some_collection_name")
# $n
# [1] 62- Add records from a third register (ISRCTN) into the same collection
Search used in this example: https://www.isrctn.com/search?q=neuroblastoma
# Retrieve trials from another register:
ctrLoadQueryIntoDb(
queryterm = "https://www.isrctn.com/search?q=neuroblastoma",
con = db
)
# * Found search query from ISRCTN: q=neuroblastoma
# * Checking trials in ISRCTN...
# Retrieved overview, records of 9 trial(s) are to be downloaded (estimate: 0.2 MB)
# (1/3) Downloading trial file...
# Download status: 1 done; 0 in progress. Total size: 93.28 Kb (100%)... done!
# (2/3) Converting to NDJSON (estimate: 0.05 s)...
# (3/3) Importing records into database...
# = Imported or updated 9 trial(s)
# Updated history ("meta-info" in "some_collection_name")
# $n
# [1] 9- Add records from a fourth register (CTIS 🔔) into the same collection
Queries in the CTIS search interface can be automatically copied to the
clipboard so that a user can paste them into queryterm, see
here.
Subsequent to the relaunch of CTIS on 2024-07-18, there are more than
4700 trials publicly accessible in CTIS. See below
for how to download documents from CTIS.
# See how many trials are in CTIS publicly accessible:
ctrLoadQueryIntoDb(
queryterm = "",
register = "CTIS",
only.count = TRUE
)
# $n
# [1] 4703
# Retrieve trials from another register:
ctrLoadQueryIntoDb(
queryterm = paste0(
'https://euclinicaltrials.eu/ctis-public/search#',
'searchCriteria={"containAll":"","containAny":"neonates","containNot":""}'),
con = db
)
# * Found search query from CTIS: searchCriteria={"containAll":"","containAny":"neonates","containNot":""}
# * Checking trials in CTIS...
# (1/4) Downloading trials list, found 7 trials
# (2/4) Downloading and processing trial data... (estimate: 0.6 Mb)
# Download status: 7 done; 0 in progress. Total size: 262.58 Kb (100%)... done!
# (3/4) Importing records into database...
# (4/4) Updating with additional data: .
# = Imported 7, updated 7 record(s) on 7 trial(s)
# Updated history ("meta-info" in "some_collection_name")
# $n
# [1] 7
allFields <- dbFindFields(".*", db, sample = TRUE)
# Finding fields in database collection (sampling 5 trial records per register) . . . . . . . .
# Field names cached for this session.
length(allFields[grepl("CTIS", names(allFields))])
# [1] 618
# root field names in CTIS
ctisFields <- allFields[grepl("CTIS", names(allFields))]
ctisFields[!grepl("[.]", ctisFields)]
# CTIS CTIS CTIS CTIS
# "ageGroup" "authorizedApplication" "correctiveMeasures" "ctNumber"
# CTIS CTIS CTIS CTIS
# "ctPublicStatus" "ctPublicStatusCode" "ctrname" "ctStatus"
# CTIS CTIS CTIS CTIS
# "decisionDate" "decisionDateOverall" "documents" "events"
# CTIS CTIS CTIS CTIS
# "gender" "lastUpdated" "publishDate" "record_last_import"
# CTIS CTIS CTIS CTIS
# "recruitmentStatus" "results" "resultsFirstReceived" "sponsorType"
# CTIS CTIS CTIS CTIS
# "startDateEU" "therapeuticAreas" "totalNumberEnrolled" "trialCountries"
# CTIS CTIS
# "trialPhase" "trialRegion"
# use an alternative to dbGetFieldsIntoDf()
allData <- nodbi::docdb_query(src = db, key = db$collection, query = '{"ctrname":"CTIS"}')
# names of top-level data items
sort(names(allData))
# [1] "_id" "ageGroup" "authorizedApplication" "correctiveMeasures"
# [5] "ctNumber" "ctPublicStatus" "ctPublicStatusCode" "ctrname"
# [9] "ctStatus" "decisionDate" "decisionDateOverall" "documents"
# [13] "events" "gender" "lastUpdated" "publishDate"
# [17] "record_last_import" "recruitmentStatus" "results" "resultsFirstReceived"
# [21] "sponsorType" "startDateEU" "therapeuticAreas" "totalNumberEnrolled"
# [25] "trialCountries" "trialPhase" "trialRegion"
# use yet another alternative
oneTrial <- DBI::dbGetQuery(
db$con, paste0(
"SELECT json(json) FROM ", db$collection,
" WHERE jsonb_extract(json, '$.ctrname') == 'CTIS'",
" LIMIT 1;"))
# display full json tree
# remotes::install_github("hrbrmstr/jsonview")
if (require(jsonview)) json_tree_view(oneTrial[[1]])
# total size of object
format(object.size(allData), "MB")
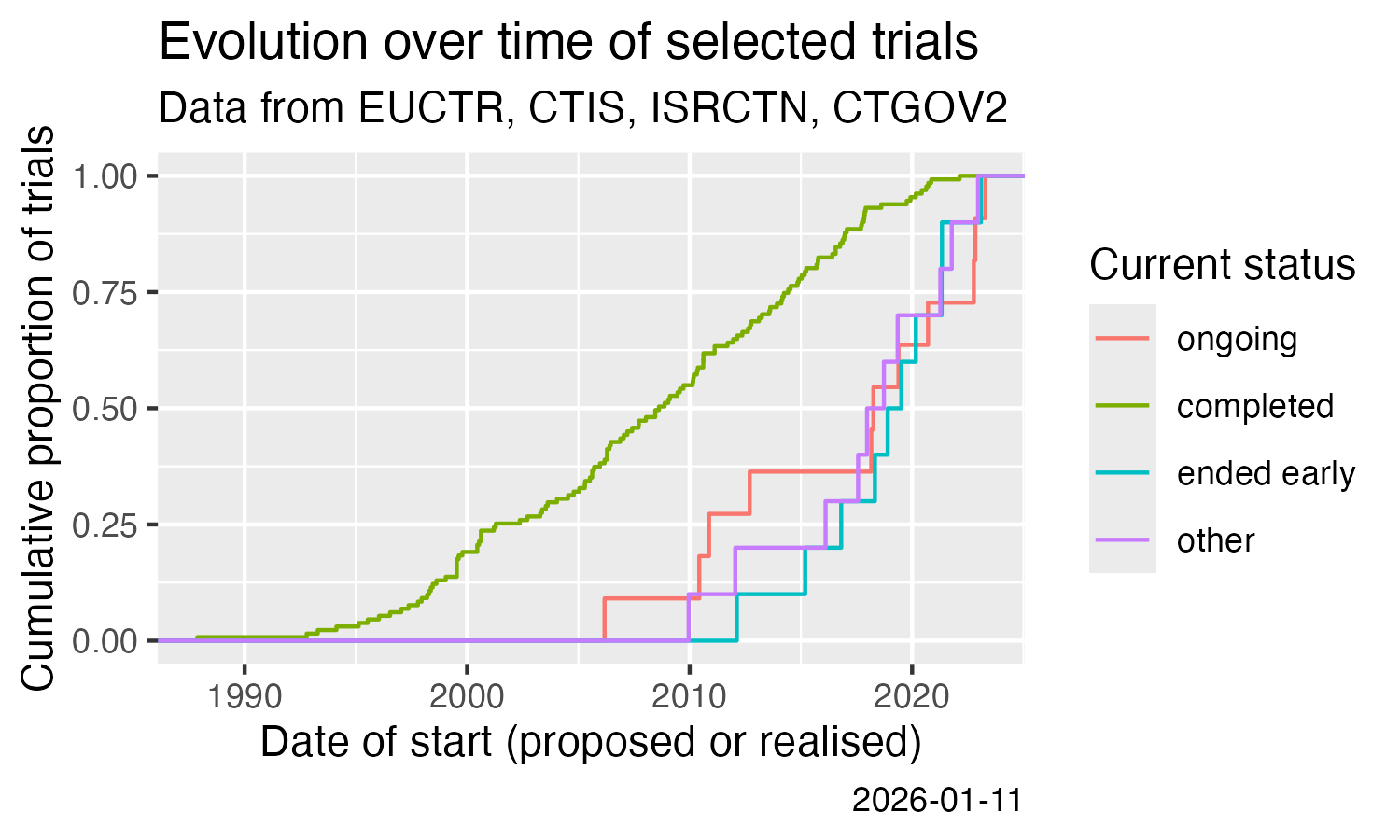
# [1] "1.4 Mb"- Analysis across trials
Show cumulative start of trials over time.
# use helper library
library(dplyr)
library(magrittr)
library(tibble)
library(purrr)
library(tidyr)
# get names of all fields / variables in the collaction
length(dbFindFields(".*", con = db))
# [1] 1667
dbFindFields("start.*date|date.*decision", con = db)
# Using cache of fields.
# - Get trial data
result <- dbGetFieldsIntoDf(
fields = c(
"ctrname",
"record_last_import",
# CTGOV2
"protocolSection.statusModule.startDateStruct.date",
"protocolSection.statusModule.overallStatus",
# EUCTR
"n_date_of_competent_authority_decision",
"trialInformation.recruitmentStartDate", # needs above: 'euctrresults = TRUE'
"p_end_of_trial_status",
# ISRCTN
"trialDesign.overallStartDate",
"trialDesign.overallEndDate",
# CTIS
"authorizedPartI.trialDetails.trialInformation.trialDuration.estimatedRecruitmentStartDate",
"ctStatus"
),
con = db
)
# - Deduplicate trials and obtain unique identifiers
# for trials that have records in several registers
# - Calculate trial start date
# - Calculate simple status for ISRCTN
# - Update end of trial status for EUCTR
result %<>%
filter(`_id` %in% dbFindIdsUniqueTrials(con = db)) %>%
rowwise() %>%
mutate(start = max(c_across(matches("(date.*decision)|(start.*date)")), na.rm = TRUE)) %>%
mutate(isrctnStatus = if_else(trialDesign.overallEndDate < record_last_import, "Ongoing", "Completed")) %>%
mutate(p_end_of_trial_status = if_else(
is.na(p_end_of_trial_status) & !is.na(n_date_of_competent_authority_decision), "Ongoing", p_end_of_trial_status)) %>%
ungroup()
# - Merge fields from different registers with re-leveling
statusValues <- list(
"ongoing" = c(
# EUCTR
"Recruiting", "Active", "Ongoing",
"Temporarily Halted", "Restarted",
# CTGOV
"Active, not recruiting", "Enrolling by invitation",
"Not yet recruiting", "ACTIVE_NOT_RECRUITING",
# CTIS
"Ongoing, recruiting", "Ongoing, recruitment ended",
"Ongoing, not yet recruiting", "Authorised, not started"
),
"completed" = c(
"Completed", "COMPLETED", "Ended"),
"other" = c(
"GB - no longer in EU/EEA", "Trial now transitioned",
"Withdrawn", "Suspended", "No longer available",
"Terminated", "TERMINATED", "Prematurely Ended",
"Under evaluation")
)
result[["state"]] <- dfMergeVariablesRelevel(
df = result,
colnames = c(
"p_end_of_trial_status",
"protocolSection.statusModule.overallStatus",
"ctStatus", "isrctnStatus"
),
levelslist = statusValues
)
# - Plot example
library(ggplot2)
ggplot(result) +
stat_ecdf(aes(x = start, colour = state)) +
labs(
title = "Evolution over time of a set of trials",
subtitle = "Data from EUCTR, CTIS, ISRCTN, CTGOV2",
x = "Date of start (proposed or realised)",
y = "Cumulative proportion of trials",
colour = "Current status",
caption = Sys.Date()
)
ggsave(
filename = "man/figures/README-ctrdata_across_registers.png",
width = 5, height = 3, units = "in"
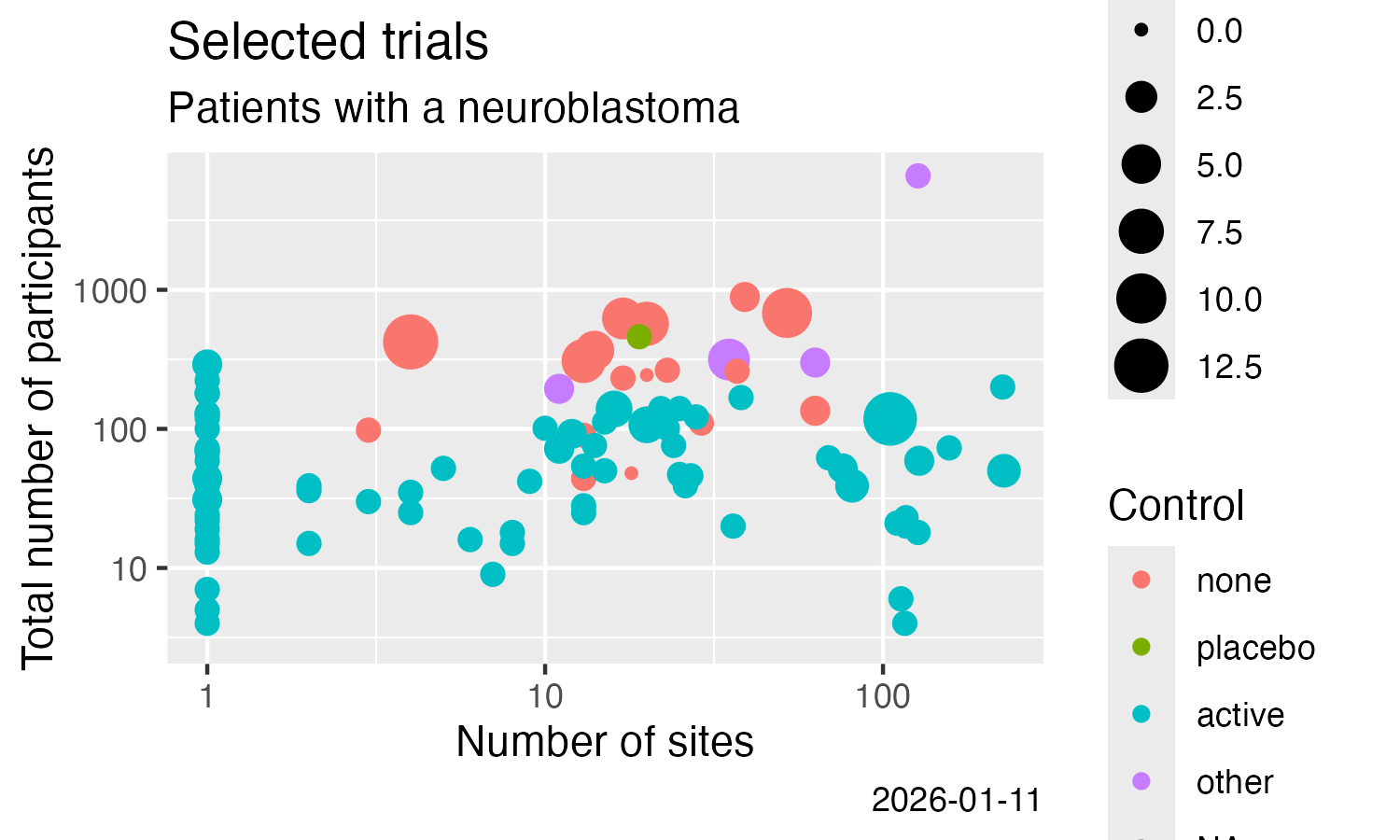
)- Result-related trial information
Analyse some simple result details, here from CTGOV2 (see this vignette for more examples):
# Get all records that have values in any of the specified fields:
result <- dbGetFieldsIntoDf(
fields = c(
# fields from CTGOV2 only
"resultsSection.baselineCharacteristicsModule.denoms.counts.value",
"resultsSection.baselineCharacteristicsModule.denoms.units",
"resultsSection.baselineCharacteristicsModule.groups.title",
"protocolSection.armsInterventionsModule.armGroups.type",
"protocolSection.designModule.designInfo.allocation",
"protocolSection.contactsLocationsModule.locations.city",
"protocolSection.conditionsModule.conditions"
),
con = db
)
# Mangle to calculate:
# - which columns with values for group counts are not labelled Total
# - what are the numbers in each of the groups etc.
result %<>%
rowwise() %>%
mutate(
number_of_arms = stringi::stri_count_fixed(
resultsSection.baselineCharacteristicsModule.groups.title, " / "),
is_randomised = case_when(
protocolSection.designModule.designInfo.allocation == "RANDOMIZED" ~ TRUE,
protocolSection.designModule.designInfo.allocation == "NON_RANDOMIZED" ~ FALSE,
number_of_arms == 1L ~ FALSE,
.default = FALSE
),
which_not_total = list(which(strsplit(
resultsSection.baselineCharacteristicsModule.groups.title, " / ")[[1]] != "Total")),
num_sites = length(strsplit(protocolSection.contactsLocationsModule.locations.city, " / ")[[1]]),
num_participants = sum(as.integer(
resultsSection.baselineCharacteristicsModule.denoms.counts.value[which_not_total])),
num_arms_or_groups = max(number_of_arms, length(which_not_total))
)
# Example plot:
library(ggplot2)
ggplot(data = result) +
labs(
title = "Trials including patients with a neuroblastoma",
subtitle = "ClinicalTrials.Gov, trials with results"
) +
geom_point(
mapping = aes(
x = num_sites,
y = num_participants,
size = num_arms_or_groups,
colour = is_randomised
)
) +
scale_x_log10() +
scale_y_log10() +
labs(
x = "Number of sites",
y = "Total number of participants",
colour = "Randomised?",
size = "# Arms / groups",
caption = Sys.Date()
)
ggsave(
filename = "man/figures/README-ctrdata_results_neuroblastoma.png",
width = 5, height = 3, units = "in"
)- Download documents: retrieve protocols, statistical analysis plans and
other documents into the local folder
./files-.../
### EUCTR document files can be downloaded when results are requested
# All files are downloaded and saved (documents.regexp is not used with EUCTR)
ctrLoadQueryIntoDb(
queryterm = "query=cancer&age=under-18&phase=phase-one",
register = "EUCTR",
euctrresults = TRUE,
documents.path = "./files-euctr/",
con = db
)
# * Found search query from EUCTR: query=cancer&age=under-18&phase=phase-one
# [...]
# Created directory ./files-euctr/
# Downloading trials...
# [...]
# = Imported or updated results for 121 trials
# = documents saved in './files-euctr'
### CTGOV files are downloaded, here corresponding to the default of
# documents.regexp = "prot|sample|statist|sap_|p1ar|p2ars|ctalett|lay|^[0-9]+ "
ctrLoadQueryIntoDb(
queryterm = "cond=Neuroblastoma&type=Intr&recrs=e&phase=1&u_prot=Y&u_sap=Y&u_icf=Y",
register = "CTGOV",
documents.path = "./files-ctgov/",
con = db
)
# * Appears specific for CTGOV Classic website
# Since 2024-06-25, the classic CTGOV servers are no longer available. Package
# ctrdata has translated the classic CTGOV query URL from this call of function
# ctrLoadQueryIntoDb(queryterm = ...) into a query URL that works with the current
# CTGOV2. This is printed below and is also part of the return value of this function,
# ctrLoadQueryIntoDb(...)$url. This URL can be used with ctrdata functions. Note that
# the fields and data schema of trials differ between CTGOV and CTGOV2.
#
# Replace this URL:
#
# https://classic.clinicaltrials.gov/ct2/results?cond=Neuroblastoma&type=Intr&recrs=
# &phase=1&u_prot=Y&u_sap=Y&u_icf=Y
#
# with this URL:
#
# https://clinicaltrials.gov/search?cond=Neuroblastoma&aggFilters=phase:2,
# docs:prot sap icf,studyType:int,status:com
#
# * Found search query from CTGOV2: cond=Neuroblastoma&aggFilters=phase:2,
# docs:prot sap icf,studyType:int,status:com
# * Checking trials using CTGOV REST API 2.0, found 26 trials
# [...]
# * Checking for documents...
# - Getting links to documents
# - Downloading documents into 'documents.path' = ./files-ctgov/
# - Created directory ./files-ctgov
# - Creating subfolder for each trial
# - Applying 'documents.regexp' to 34 documents
# - Downloading 34 missing documents
# Download status: 34 done; 0 in progress. Total size: 69.68 Mb (100%)... done!
# = Newly saved 34 document(s) for 26 trial(s); 0 document(s) for 0 trial(s)
# already existed in ./files-ctgov
### CTGOV2 files are downloaded, using the default of documents.regexp
ctrLoadQueryIntoDb(
queryterm = "https://clinicaltrials.gov/search?cond=neuroblastoma&aggFilters=phase:1,results:with",
documents.path = "./files-ctgov2/",
con = db
)
# * Found search query from CTGOV2: cond=neuroblastoma&aggFilters=phase:1,results:with
# [...]
# * Checking for documents...
# - Getting links to documents
# - Downloading documents into 'documents.path' = ./files-ctgov2/
# - Created directory ./files-ctgov2
# - Creating subfolder for each trial
# - Applying 'documents.regexp' to 35 documents
# - Downloading 35 missing documents
# Download status: 35 done; 0 in progress. Total size: 76.64 Mb (100%)... done!
# = Newly saved 35 document(s) for 22 trial(s); 0 document(s) for 0 trial(s) already
# existed in ./files-ctgov2
### ISRCTN files are downloaded, using the default of documents.regexp
ctrLoadQueryIntoDb(
queryterm = "https://www.isrctn.com/search?q=alzheimer",
documents.path = "./files-isrctn/",
con = db
)
# * Found search query from ISRCTN: q=alzheimer
# [...]
# * Checking for documents...
# - Getting links to documents
# - Downloading documents into 'documents.path' = ./files-isrctn/
# - Created directory ./files-isrctn
# - Creating subfolder for each trial
# - Applying 'documents.regexp' to 41 documents
# - Downloading 26 missing documents
# Download status: 26 done; 0 in progress. Total size: 12.83 Mb (100%)... done!
# Download status: 2 done; 0 in progress. Total size: 6.56 Kb (100%)... done!
# = Newly saved 24 document(s) for 12 trial(s); 0 document(s) for 0 trial(s)
# already existed in ./files-isrctn
### CTIS files are downloaded, using the default of documents.regexp
ctrLoadQueryIntoDb(
queryterm = paste0(
'https://euclinicaltrials.eu/ctis-public/search#',
'searchCriteria={"containAll":"","containAny":"cancer","containNot":""}'),
documents.path = "./files-ctis/",
con = db
)
# * Found search query from CTIS: searchCriteria={"containAll":"","containAny":"cancer","containNot":""}
# * Checking trials in CTIS...
# (1/4) Downloading trial list(s), found 1109 trials
# (2/4) Downloading and processing trial data... (estimate: 100 Mb)
# (3/4) Importing records into database...
# (4/4) Updating with additional data: .
# * Checking for documents...
# - Downloading documents into 'documents.path' = ./files-ctis/
# - Created directory ./files-ctis
# - Creating subfolder for each trial
# - Applying 'documents.regexp' to 20 documents
# - Downloading 13 missing documents
# Download status: 13 done; 0 in progress. Total size: 8.07 Mb (100%)... done!
# = Newly saved 13 document(s) for 4 trial(s); 0 document(s) for 0 trial(s)
# already existed in ./files-ctis
# = Imported 1109, updated 1109 record(s) on 1109 trial(s)
# No history found in expected format.
# Updated history ("meta-info" in "some_collection_name")
# $n
# [1] 1109See also https://app.codecov.io/gh/rfhb/ctrdata/tree/master/R
tinytest::test_all()
# test_ctrdata_ctrfindactivesubstance.R 4 tests OK 1.6s
# test_ctrdata_duckdb_ctgov2.R.. 50 tests OK 2.4s
# test_ctrdata_duckdb_ctis.R.... 172 tests OK 15.2s
# test_ctrdata_mongo_local_ctgov.R 51 tests OK 57.7s
# test_ctrdata_other_functions.R 64 tests OK 3.8s
# test_ctrdata_postgres_ctgov2.R 50 tests OK 2.6s
# test_ctrdata_sqlite_ctgov.R... 52 tests OK 56.0s
# test_ctrdata_sqlite_ctgov2.R.. 50 tests OK 2.3s
# test_ctrdata_sqlite_ctis.R.... 194 tests OK 12.5s
# test_ctrdata_sqlite_euctr.R... 105 tests OK 1.3s
# test_ctrdata_sqlite_isrctn.R.. 38 tests OK 21.4s
# test_euctr_error_sample.R..... 8 tests OK 0.9s
# All ok, 838 results (38m 48.8s)
covr::package_coverage(path = ".", type = "tests")
# ctrdata Coverage: 93.68%
# R/zzz.R: 80.95%
# R/ctrRerunQuery.R: 89.16%
# R/ctrLoadQueryIntoDbEuctr.R: 90.03%
# R/utils.R: 90.89%
# R/ctrLoadQueryIntoDbIsrctn.R: 92.11%
# R/dbGetFieldsIntoDf.R: 93.06%
# R/ctrLoadQueryIntoDbCtgov2.R: 94.05%
# R/ctrLoadQueryIntoDb.R: 94.12%
# R/ctrLoadQueryIntoDbCtis.R: 94.13%
# R/ctrLoadQueryIntoDbCtgov.R: 95.04%
# R/dbFindFields.R: 95.24%
# R/ctrGetQueryUrl.R: 96.00%
# R/ctrOpenSearchPagesInBrowser.R: 97.22%
# R/dfMergeVariablesRelevel.R: 97.30%
# R/dfTrials2Long.R: 97.35%
# R/dbFindIdsUniqueTrials.R: 97.77%
# R/dfName2Value.R: 98.61%
# R/ctrFindActiveSubstanceSynonyms.R: 100.00%
# R/dbQueryHistory.R: 100.00%-
See project outline https://github.com/users/rfhb/projects/1
-
Canonical definitions, filters, calculations are in the works (since August 2023) for data mangling and analyses across registers, e.g. to define study population, identify interventional trials, calculate study duration; public collaboration on these canonical scripts will speed up harmonising analyses.
-
Merge results-related fields retrieved from different registers, such as corresponding endpoints (work not yet started). The challenge is the incomplete congruency and different structure of data fields.
-
Authentication, expected to be required by CTGOV2; specifications not yet known (work not yet started).
-
Explore further registers (exploration is continually ongoing; added value, terms and conditions for programmatic access vary; no clear roadmap is established yet).
-
Retrieve previous versions of protocol- or results-related information. The challenges include, historic versions can only be retrieved one-by-one, do not include results, or are not in structured format.(functionality available with version 1.17.2.9000 to the extent that seems reasonably possible at this time, namely for protocol-related information for CTIS and for protocol- and results-related information in CTGOV2)
-
Data providers and curators of the clinical trial registers. Please review and respect their copyrights and terms and conditions, see
ctrOpenSearchPagesInBrowser(copyright = TRUE). -
Package
ctrdatahas been made possible building on the work done for R, clipr. curl, dplyr, duckdb, httr, jqr, jsonlite, lubridate, mongolite, nodbi, RPostgres, RSQLite, rvest, stringi and xml2.
-
Please file issues and bugs here. Also check out how to handle some of the closed issues, e.g. on C stack usage too close to the limit and on a SSL certificate problem: unable to get local issuer certificate
-
Information in trial registers may not be fully correct; see for example this publication on CTGOV.
-
No attempts were made to harmonise field names between registers (nevertheless,
dfMergeVariablesRelevel()can be used to merge and map several variables / fields into one).