holoviz / hvplot Goto Github PK
View Code? Open in Web Editor NEWA high-level plotting API for pandas, dask, xarray, and networkx built on HoloViews
Home Page: https://hvplot.holoviz.org
License: BSD 3-Clause "New" or "Revised" License
A high-level plotting API for pandas, dask, xarray, and networkx built on HoloViews
Home Page: https://hvplot.holoviz.org
License: BSD 3-Clause "New" or "Revised" License
import pandas as pd
import numpy as np
import dask.dataframe as dd
import holoplot.dask
import holoplot.pandas
def generate_df(size):
d = {
'dates' : pd.date_range('1980-01-01', periods=size, freq='1T'),
'unique_id' : np.arange(0, size),
'ints' : np.random.randint(0, size, size=size),
'floats' : np.random.randn(size),
'bools' : np.random.choice([0, 1], size=size),
'int_nans' : np.random.choice([0, 1, np.nan], size=size),
'float_nans' : np.random.choice([0.0, 1.0, np.nan], size=size),
'constant' : 1,
'fake_categorical' : np.random.choice([10, 20, 30, 40, 50], size=size) ,
'categorical_binary' : np.random.choice(['a', 'b'], size=size),
'categorical_nans' : np.random.choice(['a', 'b', 'c', 'd', 'e', 'f', 'g'], size=size),
'signal1' : np.random.normal(0, 0.3, size=size).cumsum() + 50,
'signal2' : np.random.normal(0, 0.3, size=size).cumsum() + 50
}
df = pd.DataFrame(d)
df['hardbools'] = df['bools'] == 1
df['categorical_nans'] = df['categorical_nans'].replace('c', np.nan)
df['categorical_binary'] = df['categorical_binary'].astype('category')
df['unique_id'] = df['unique_id'].astype(str)
df.index = df.dates
df.index.name = 'dates'
df = df.drop('dates', axis=1)
return df
def generate_dask_df(size, npartitions=32):
df = generate_df(size)
df = dd.from_pandas(df, npartitions=npartitions)
return dfThese work fine:
df.holoplot.line(y='signal1')
df.holoplot.line(y='signal1', by='categorical_binary')
df.holoplot.line(y='signal1', datashade=True)
This is the result for by and datashade :
df.holoplot.line(y='signal1', by='categorical_binary', datashade=True)

Geographic Plots Using GeoViews and Cartopy to plot data in geographic coordinate systems. ->
Geographic Plots Using GeoViews, Cartopy, and Geopandas to plot data in geographic coordinate systems.
import os
import xarray as xr
import holoviews as hv
import geoviews as gv
import holoplot.xarray
hv.extension('bokeh')
os.system(
'wget '
'ftp://ftp.cdc.noaa.gov/'
'Datasets/ncep.reanalysis.dailyavgs/'
'surface/'
'air.sig995.2018.nc'
)
obs_ds = xr.open_dataset('air.sig995.2018.nc')
obs_ds.holoplot.image(x='lon', y='lat', z='air', groupby='time')
obs_ds.holoplot.image(x='lon', y='lat', z='air', groupby='time') * gv.feature.coastline()
hv.Dataset(obs_ds).to(gv.Image, ['lon', 'lat'], 'air', ['time'], dynamic=True) * gv.feature.coastline()
shouldn't it be holoplot now ?
import os
import xarray as xr
import hvplot.xarray
os.system(
'wget -nc '
'ftp://ftp.cdc.noaa.gov/'
'Datasets/ncep.reanalysis.dailyavgs/'
'surface/'
'air.sig995.2018.nc'
)
obs_ds = xr.open_dataset('air.sig995.2018.nc')
# this works
obs_ds.hvplot.contourf(x='lon', y='lat', z='air', groupby='time')
# not this
obs_ds.hvplot.contourf(x='lon', y='lat', z='air', groupby='time', geo=True)
AxisError: axis 1 is out of bounds for array of dimension 1
I was wondering if there's any interest in integrating a built-in theme to hvplot? If not, I'm thinking of creating a separate package that, on import, applies this theme (like seaborn).
Examples of how it compares to https://pyviz.github.io/hvplot/user_guide/Gridded_Data.html
https://anaconda.org/ahuang11/hvplot_theme/notebook
and also https://pyviz.github.io/hvplot/user_guide/Introduction.html


There are a couple of issues though.
So I was thinking of wrapping this into a function with adjustable keyword arguments (and also remedy the title) that returns hv.renderer('bokeh').theme = theme
station_df.max_temp_f.hist(alpha=0.5, bins=[50, 60, 70])
vs
station_df.max_temp_f.holoplot.hist(alpha=0.5, bins=[50, 60, 70])
station_df.max_temp_f.holoplot.hist(alpha=0.5, bins=[50, 60, 70])
...
~/anaconda3/lib/python3.6/site-packages/holoviews/operation/element.py in _process(self, view, key)
556
557 datetimes = False
--> 558 steps = self.p.num_bins + 1
559 start, end = hist_range
560 if data.dtype.kind == 'M' or (data.dtype.kind == 'O' and isinstance(data[0], datetime_types)):
TypeError: can only concatenate list (not "int") to list
Currently no errorbars are supported, it should be possible to add them for Curve, Area, Bars, Scatter
Raise a warning, and also generate a generic linspace in the background for xarray objects that don't have coordinates attached so can quickly visualize the data.
<xarray.DataArray 'Field020' (Field020_dim002: 90, Field020_dim001: 180)>
array([[ 4.219483e-03, 8.433826e-03, 1.263789e-02, ..., -8.432080e-03,
-4.217734e-03, 1.750679e-06],
[ 5.359092e-03, 1.071166e-02, 1.605117e-02, ..., -1.070944e-02,
-5.356870e-03, 2.223507e-06],
[ 6.492877e-03, 1.297784e-02, 1.944700e-02, ..., -1.297516e-02,
-6.490185e-03, 2.693918e-06],
...,
[ 5.292206e-03, 1.057796e-02, 1.585083e-02, ..., -1.057577e-02,
-5.290011e-03, 2.195755e-06],
[ 4.152293e-03, 8.299527e-03, 1.243665e-02, ..., -8.297808e-03,
-4.150571e-03, 1.722801e-06],
[ 3.007867e-03, 6.012070e-03, 9.008948e-03, ..., -6.010825e-03,
-3.006620e-03, 1.247975e-06]])
Dimensions without coordinates: Field020_dim002, Field020_dim001
~/anaconda3/lib/python3.6/site-packages/hvplot/util.py in process_xarray(data, x, y, by, groupby, use_dask, persist, gridded, label, value_label)
276 x = data_vars[0] if y in dims else dims[0]
277 elif not x and not y:
--> 278 x, y = dims[0], data_vars
279
280 if by is None:
IndexError: list index out of range
Hi I am just starting with hvplot and it looks phenomenal.
I am however encountering a weird error:
When running it on jupyterlab the bokeh window 'floats' over the interface
(I am not sure how to add a screenshot of the issue...)
here is the code I use
import xarray as xr
import numpy as np
import hvplot.dask
import hvplot.xarray
import holoviews as hv
oxygen.o2.isel(time=10, st_ocean=10).hvplot()where oxygen is this dataset:
<xarray.DataArray 'o2' (time: 36, st_ocean: 50, yt_ocean: 200, xt_ocean: 360)>
dask.array<shape=(36, 50, 200, 360), dtype=float32, chunksize=(1, 50, 200, 360)>
Coordinates:
run <U7 'control'
* st_ocean (st_ocean) float64 5.034 15.1 25.22 35.36 45.58 55.85 ...
* xt_ocean (xt_ocean) float64 -279.5 -278.5 -277.5 -276.5 -275.5 ...
* yt_ocean (yt_ocean) float64 -81.5 -80.5 -79.5 -78.5 -77.5 -76.5 ...
* time (time) datetime64[ns] 2181-01-16T12:00:00 2181-02-15 ...
geolon_t (yt_ocean, xt_ocean) float64 -279.5 -278.5 -277.5 -276.5 ...
geolat_t (yt_ocean, xt_ocean) float64 -81.5 -81.5 -81.5 -81.5 -81.5 ...
area_t (yt_ocean, xt_ocean) float64 1.828e+09 1.828e+09 1.828e+09 ...
dxt (yt_ocean, xt_ocean) float64 1.644e+04 1.644e+04 1.644e+04 ...
dyt (yt_ocean, xt_ocean) float64 1.112e+05 1.112e+05 1.112e+05 ...
ht (yt_ocean, xt_ocean) float64 0.0 0.0 0.0 0.0 0.0 0.0 0.0 ...
kmt (yt_ocean, xt_ocean) float64 0.0 0.0 0.0 0.0 0.0 0.0 0.0 ...
wet_t (yt_ocean, xt_ocean) float64 0.0 0.0 0.0 0.0 0.0 0.0 0.0 ...
tmask (yt_ocean, xt_ocean) float64 0.0 0.0 0.0 0.0 0.0 0.0 0.0 ...
tmask_region (yt_ocean, xt_ocean) float64 0.0 0.0 0.0 0.0 0.0 0.0 0.0 ...
Attributes:
long_name: Oxygen (mini)
units: mol/kg
hvplot.__version__
'0.2.1'
Add to the documenation for "Getting Started", that to get the us_crime and airline_flights catalogues you have to install the packages with: conda install -c intake us_crime airline_flights. Then to load them, I had to use crime = intake.Catalog('[direct path to yaml file]'), instead of intake.cat.us_crime.read() which gave me a KeyError. Once loaded, I got the dataframe by calling crime.us_crime.read(). Similar process was used to load the flights dataframe.

import os
import xarray as xr
import holoplot.xarray
os.system('wget http://ftp.cpc.ncep.noaa.gov/NMME/realtime_anom/NCAR_CCSM4/2018050800/NCAR_CCSM4.tmpsfc.201805.anom.nc')
ds = xr.open_dataset('NCAR_CCSM4.tmpsfc.201805.anom.nc', decode_cf=False)
ds.fcst.isel(ensmem=0, target=0).plot()
ds.fcst.isel(ensmem=0, target=0).plot(x='lon', y='lat')
Allow overriding the axis label easily using xlabel/ylabel arguments.
obs_ds.holoplot.image('lon', 'lat').options(projection=crs.Robinson())
world = gpd.read_file(gpd.datasets.get_path('naturalearth_lowres'))
world.hvplot(width=550) + world.hvplot(groupby='continent', width=500)
KeyError: "Dimension 'continent' not found."
Still points to holoplot docs site, which no longer exists
I thought from holoviz/holoviews#30 (comment) that this might work:
import xarray as xr
import hvplot.xarray
url = 'https://rsignell.s3.amazonaws.com/NLCD/nlcd_2011_landcover_2011_edition_2014_10_10.tif'
da = xr.open_rasterio(url, chunks={'x':2056, 'y':2056})
%%opts Image [width=700 height=500 colorbar=True aspect='equal'] (cmap='magma')
da[0, 50000:51000, 80000:81000].hvplot()but I guess aspect is not a thing.
Is it possible to use Raster with hvplot?
I couldn't figure that out either. 😿
If trying to holoplot a 2D xr.DataArray, it currently must have a name to work. These names are usually inherited when e.g. selecting from a xr.Dataset but are seemingly dropped after some operations. A quick fix would just be an informative error.
Some code to reproduce:
import numpy as np
import xarray as xr
import holoplot.xarray
da = xr.DataArray(
data=np.random.randn(10, 10),
dims=['a', 'b'],
coords={'a': np.arange(10), 'b': np.arange(10)},
)
da.holoplot.image()produces the slightly obscure error:
...
~/conda/lib/python3.6/site-packages/holoplot/converter.py in image(self, x, y, z, data)
859 element = self._get_element('image')
860 if self.geo: params['crs'] = self.crs
--> 861 return element(data, [x, y], z, **params).redim(**self._redim).redim.range(**ranges).opts(**opts)
862
863 def quadmesh(self, x=None, y=None, z=None, data=None):
TypeError: range() keywords must be stringswhereas with a name set:
da = xr.DataArray(
data=np.random.randn(10, 10),
dims=['a', 'b'],
coords={'a': np.arange(10), 'b': np.arange(10)},
name='test',
)
da.holoplot.image()It would be really helpful if hvplot for intake allowed the user to subset the data in the catalog for declarative plotting. https://intake.readthedocs.io/en/latest/plotting.html#declaring-defaults.
Maybe something like:
select:
- latitude: [-45, 45]To me, typing holoplot in its entirety can be a bit tedious.
The hvplot namespace is polluted by param methods and attributes which I don't think are needed which means that tab-completing the plot methods doesn't work well. Additionally we should provide an easy way of listing the available plot methods.
This issue is a follow up on this SO answer as per @philippjfr request. Please keep in mind it's the first time I'm using holoplot (which seems to be awesome!)
I tried to use holoplot from within a Jupyter notebook. Once importing the package using
import holoplot.pandasall .plot are going through holoplot. This is not always a convenient behavior. Sometimes, it is easy to think of a use case where some cells should be using holoplot while others will be rendered using the standard matplotlib. I hope this issue is good enough as a starting point for an interesting discussion.
import holoviews as hv
hv.extension('bokeh')
import geoviews
from shapely.geometry import Point
import pandas as pd
import geopandas as gpd
from geopandas import GeoSeries, GeoDataFrame
import hvplot.pandas
from geopandas import GeoSeries, GeoDataFrame
from shapely.geometry import Point
data = {'name': ['House', 'Work', 'Pet Store'],
'lat': [ 45, 46, 47.5],
'lon': [-120, -121.2, -122.9]}
df = pd.DataFrame(data)
geo = [Point(xy) for xy in zip(data['lon'], data['lat'])]
gdf = GeoDataFrame(df, geometry=geo)
gdf.hvplot().options(size=15,color='green',marker='*')
The plot correctly sizes the axis ranges,
but the third point is not displayed
{'matplotlib': '2.2.2', 'pandas': '0.23.2', 'geopandas': '0.3.0', 'holoviews': '1.11.0a1.post3+g103daf59-dirty'}
hvplot from master v0.2.1 (july 9)
df contains three columns
df.holoplot(color=hv.Cycle(['red', 'blue', 'black']))
/mnt/c/Users/sephi/GOOGLE~1/Bash/holoplot/holoplot/__init__.py in __call__(self, x, y, kind, **kwds)
76 kind = kind or params.pop('kind', None)
77 converter = HoloViewsConverter(
---> 78 self._data, x, y, kind=kind, **params
79 )
80 return converter(kind, x, y)
/mnt/c/Users/sephi/GOOGLE~1/Bash/holoplot/holoplot/converter.py in __init__(self, data, x, y, kind, by, use_index, group_label, value_label, backlog, persist, use_dask, crs, fields, groupby, dynamic, width, height, shared_axes, grid, legend, rot, title, xlim, ylim, clim, xticks, yticks, logx, logy, loglog, hover, subplots, label, invert, stacked, colorbar, fontsize, colormap, datashade, rasterize, row, col, figsize, debug, xaxis, yaxis, framewise, aggregator, projection, global_extent, geo, **kwds)
206 elif not isinstance(color, (Cycle, list)):
207 continue
--> 208 style_opts[opt] = Cycle(values=cycle) if isinstance(cycle, list) else cycle
209 self._style_opts = style_opts
210 self._norm_opts = {'framewise': framewise, 'axiswise': not shared_axes}
UnboundLocalError: local variable 'cycle' referenced before assignment
I get an error trying introduction.ipynb, did I miss something ?
import holoplot.pandas
df.holoplot()
---------------------------------------------------------------------------
AttributeError Traceback (most recent call last)
<ipython-input-3-5a136e144f5a> in <module>()
1 import holoplot.pandas
2
----> 3 df.holoplot()
C:\WinP\bd36\buQt5\winp32-3.6.x.1\python-3.6.5\lib\site-packages\pandas\core\generic.py in __getattr__(self, name)
4370 if self._info_axis._can_hold_identifiers_and_holds_name(name):
4371 return self[name]
-> 4372 return object.__getattribute__(self, name)
4373
4374 def __setattr__(self, name, value):
AttributeError: 'DataFrame' object has no attribute 'holoplot'
This came out of a conversation with @philippjfr. Potentially crs could also accept a string which would refer to an attr or I guess a coord containing the crs info. This would be a little trickier to implement since crs already accepts a proj4 string. But I think it'd be nicely parallel to how x, y, and z are set.
When I do something like
%%opts BoxWhisker {-framewise}
from bokeh.sampledata.autompg import autompg as df
title = "MPG by Cylinders and Data Source, Colored by Cylinders"
boxwhisker = hv.BoxWhisker(df, ['cyl', 'origin'], 'mpg', label=title)
df.plot.box('mpg', 'origin', groupby='yr')
the ylimits do not stay consistent across frames.

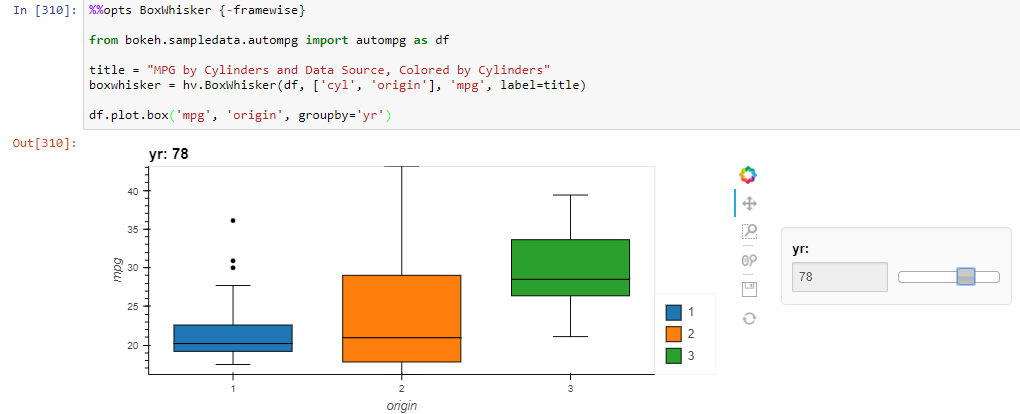
Works with HoloViews dataset.to.box syntax though
%%opts BoxWhisker {-framewise}
from bokeh.sampledata.autompg import autompg as df
title = "MPG by Cylinders and Data Source, Colored by Cylinders"
boxwhisker = hv.BoxWhisker(df, ['cyl', 'origin'], 'mpg', label=title)
hv.Dataset(df).to.box('origin', 'mpg', groupby='yr')
geopandas.read_file('LondonBoroughs.shp').hvplot('PopDen').options(cmap='RdBu', colorbar=True)
returns an hv.Polygon element without the colorbar
tst = pd.DataFrame(data={'lon': [1, 2, 3], 'lat': [4, 3, 2], 'idx': [0, 3, 5]})
tst.hvplot('lon', 'lat', geo=True, framewise=False).options(line_width=75) \
* tst.hvplot.points('lon', 'lat', groupby='idx', geo=True, framewise=False)
This yields unexpected results (the points aren't overlaid on top of curve).
A bonus would be to replace hv.Curve with gv.Path if geo=True.
results in the following stacktrace
---------------------------------------------------------------------------
TypeError Traceback (most recent call last)
<ipython-input-10-02b67fc4e6b3> in <module>()
----> 1 c.fuzz_udp.plot.hist()
/opt/conda/lib/python3.6/site-packages/holoplot/__init__.py in hist(self, y, by, **kwds)
300 The HoloViews representation of the plot.
301 """
--> 302 return self(kind='hist', x=None, y=y, by=by, **kwds)
303
304 def kde(self, y=None, by=None, **kwds):
/opt/conda/lib/python3.6/site-packages/holoplot/__init__.py in __call__(self, x, y, kind, **kwds)
81 self._data, x, y, kind=kind, **params
82 )
---> 83 return converter(kind, x, y)
84
85 def line(self, x=None, y=None, **kwds):
/opt/conda/lib/python3.6/site-packages/holoplot/converter.py in __call__(self, kind, x, y)
520 obj = DynamicMap(cbcallable, streams=[self.stream])
521 else:
--> 522 obj = method(x, y)
523
524 if not (self.datashade or self.rasterize):
/opt/conda/lib/python3.6/site-packages/holoplot/converter.py in hist(self, x, y, data)
760 hists = []
761 for col in y:
--> 762 hist = histogram(ds, dimension=col, **hist_opts)
763 ranges = {hist.kdims[0].name: self._dim_ranges['x'],
764 hist.vdims[0].name: self._dim_ranges['y']}
/opt/conda/lib/python3.6/site-packages/param/parameterized.py in __new__(class_, *args, **params)
1889 inst = class_.instance()
1890 inst._set_name(class_.__name__)
-> 1891 return inst.__call__(*args,**params)
1892
1893 def __call__(self,*args,**kw):
/opt/conda/lib/python3.6/site-packages/holoviews/core/operation.py in __call__(self, element, **params)
161 operation=self, kwargs=params)
162 elif isinstance(element, ViewableElement):
--> 163 processed = self._apply(element)
164 elif isinstance(element, DynamicMap):
165 if any((not d.values) for d in element.kdims):
/opt/conda/lib/python3.6/site-packages/holoviews/core/operation.py in _apply(self, element, key)
119 for hook in self._preprocess_hooks:
120 kwargs.update(hook(self, element))
--> 121 ret = self._process(element, key)
122 for hook in self._postprocess_hooks:
123 ret = hook(self, ret, **kwargs)
/opt/conda/lib/python3.6/site-packages/holoviews/operation/element.py in _process(self, view, key)
549 weights = None
550
--> 551 data = data[np.isfinite(data)]
552 hist_range = self.p.bin_range or view.range(selected_dim)
553 # Avoids range issues including zero bin range and empty bins
TypeError: ufunc 'isfinite' not supported for the input types, and the inputs could not be safely coerced to any supported types according to the casting rule ''safe'
Currently the hover tool shows only the x/y columns and if enabled the color columns but you should be able to add additional columns.
Similar to hv.Dataset(df).to.labels()
Hi all, I'm trying to see if using holoplot is feasible for an upcoming talk on real-time processing, for which streamz will make a brief appearance. I'm currently not getting much.
Is there additional setup that I should be doing here? Is there something that I can do to accelerate tracking down issues here?
import os
import xarray as xr
import holoviews as hv
import geoviews as gv
import holoplot.xarray
hv.extension('bokeh')
os.system(
'wget '
'ftp://ftp.cdc.noaa.gov/'
'Datasets/ncep.reanalysis.dailyavgs/'
'surface/'
'air.sig995.2018.nc'
)
obs_ds = xr.open_dataset('air.sig995.2018.nc')
obs_ds.holoplot.quadmesh(x='lon', y='lat', z='air', groupby='time')
obs_ds.holoplot.image(x='lon', y='lat', z='air', groupby='time')
Or maybe I just don't really know what the difference between quadmesh and image is.
tst = (hv.Curve([0, 1, 2], group='agroup', label='alabel') + hv.Curve([3, 4, 5], group='bgroup', label='blabel'))
print(tst)
print('\nhvplot')
tst_df = pd.DataFrame({'x1': [0, 1, 2], 'x2': [3, 4, 5]})
print(tst_df.hvplot('index', 'x1', label='alabel', group_label='agroup') + tst_df.hvplot('index', 'x2', label='blabel', group_label='bgroup'))
I've successfully used hvplot to make super-quick maps of some global wave model data with time slider as described in the gridded data user guide:

But for other global wave model data, I still get the histogram using the same approach:

The notebook is here:
https://nbviewer.jupyter.org/gist/rsignell-usgs/3ff85b150bfd118f2b2ed768f98116a1
Any idea why and how I can work around this?
The following examples gives an error.
If you compute() dataframe it will work.
If it is non datashaded, it will work.
def generate_df(size):
d = {
'dates' : pd.date_range('1980-01-01', periods=size, freq='1T'),
'unique_id' : np.arange(0, size),
'ints' : np.random.randint(0, size, size=size),
'floats' : np.random.randn(size),
'bools' : np.random.choice([0, 1], size=size),
'int_nans' : np.random.choice([0, 1, np.nan], size=size),
'float_nans' : np.random.choice([0.0, 1.0, np.nan], size=size),
'constant' : 1,
'fake_categorical' : np.random.choice([10, 20, 30, 40, 50], size=size) ,
'categorical_binary' : np.random.choice(['a', 'b'], size=size),
'categorical_nans' : np.random.choice(['a', 'b', 'c', 'd', 'e', 'f', 'g'], size=size),
'signal1' : np.random.normal(0, 0.3, size=size).cumsum() + 50,
'signal2' : np.random.normal(0, 0.3, size=size).cumsum() + 50
}
df = pd.DataFrame(d)
df['hardbools'] = df['bools'] == 1
df['categorical_nans'] = df['categorical_nans'].replace('c', np.nan)
df['categorical_binary'] = df['categorical_binary'].astype('category')
df['unique_id'] = df['unique_id'].astype(str)
df.index = df.dates
df.index.name = 'dates'
df = df.drop('dates', axis=1)
return df
def generate_dask_df(size, npartitions=16):
return dd.from_pandas(generate_df(size), npartitions=npartitions)
import holoplot.dask
import holoplot.pandas
plt = df.holoplot(y='signal1', by="categorical_binary",
title='yo', datashade=True,
)
plt
This gives stacktrace:
KeyError Traceback (most recent call last)
~/jungle/local/lib/python3.5/site-packages/ipython-6.3.1-py3.5.egg/IPython/core/formatters.py in __call__(self, obj, include, exclude)
968
969 if method is not None:
--> 970 return method(include=include, exclude=exclude)
971 return None
972 else:
~/area51/holoviews/holoviews/core/dimension.py in _repr_mimebundle_(self, include, exclude)
1229 combined and returned.
1230 """
-> 1231 return Store.render(self)
1232
1233
~/area51/holoviews/holoviews/core/options.py in render(cls, obj)
1287 data, metadata = {}, {}
1288 for hook in hooks:
-> 1289 ret = hook(obj)
1290 if ret is None:
1291 continue
~/area51/holoviews/holoviews/ipython/display_hooks.py in pprint_display(obj)
270 if not ip.display_formatter.formatters['text/plain'].pprint:
271 return None
--> 272 return display(obj, raw_output=True)
273
274
~/area51/holoviews/holoviews/ipython/display_hooks.py in display(obj, raw_output, **kwargs)
246 elif isinstance(obj, (HoloMap, DynamicMap)):
247 with option_state(obj):
--> 248 output = map_display(obj)
249 elif isinstance(obj, Plot):
250 output = render(obj)
~/area51/holoviews/holoviews/ipython/display_hooks.py in wrapped(element)
140 try:
141 max_frames = OutputSettings.options['max_frames']
--> 142 mimebundle = fn(element, max_frames=max_frames)
143 if mimebundle is None:
144 return {}, {}
~/area51/holoviews/holoviews/ipython/display_hooks.py in map_display(vmap, max_frames)
200 return None
201
--> 202 return render(vmap)
203
204
~/area51/holoviews/holoviews/ipython/display_hooks.py in render(obj, **kwargs)
63 renderer = renderer.instance(fig='png')
64
---> 65 return renderer.components(obj, **kwargs)
66
67
~/area51/holoviews/holoviews/plotting/bokeh/renderer.py in components(self, obj, fmt, comm, **kwargs)
260 # Bokeh has to handle comms directly in <0.12.15
261 comm = False if bokeh_version < '0.12.15' else comm
--> 262 return super(BokehRenderer, self).components(obj,fmt, comm, **kwargs)
263
264
~/area51/holoviews/holoviews/plotting/renderer.py in components(self, obj, fmt, comm, **kwargs)
327 plot = obj
328 else:
--> 329 plot, fmt = self._validate(obj, fmt)
330
331 widget_id = None
~/area51/holoviews/holoviews/plotting/renderer.py in _validate(self, obj, fmt, **kwargs)
226 if isinstance(obj, tuple(self.widgets.values())):
227 return obj, 'html'
--> 228 plot = self.get_plot(obj, renderer=self, **kwargs)
229
230 fig_formats = self.mode_formats['fig'][self.mode]
~/area51/holoviews/holoviews/plotting/bokeh/renderer.py in get_plot(self_or_cls, obj, doc, renderer)
153 curdoc().theme = self_or_cls.theme
154 doc.theme = self_or_cls.theme
--> 155 plot = super(BokehRenderer, self_or_cls).get_plot(obj, renderer)
156 plot.document = doc
157 return plot
~/area51/holoviews/holoviews/plotting/renderer.py in get_plot(self_or_cls, obj, renderer)
193
194 # Initialize DynamicMaps with first data item
--> 195 initialize_dynamic(obj)
196
197 if not isinstance(obj, Plot):
~/area51/holoviews/holoviews/plotting/util.py in initialize_dynamic(obj)
242 continue
243 if not len(dmap):
--> 244 dmap[dmap._initial_key()]
245
246
~/area51/holoviews/holoviews/core/spaces.py in __getitem__(self, key)
1130 # Not a cross product and nothing cached so compute element.
1131 if cache is not None: return cache
-> 1132 val = self._execute_callback(*tuple_key)
1133 if data_slice:
1134 val = self._dataslice(val, data_slice)
~/area51/holoviews/holoviews/core/spaces.py in _execute_callback(self, *args)
909
910 with dynamicmap_memoization(self.callback, self.streams):
--> 911 retval = self.callback(*args, **kwargs)
912 return self._style(retval)
913
~/area51/holoviews/holoviews/core/spaces.py in __call__(self, *args, **kwargs)
570
571 try:
--> 572 ret = self.callable(*args, **kwargs)
573 except KeyError:
574 # KeyError is caught separately because it is used to signal
~/area51/holoviews/holoviews/util/__init__.py in dynamic_operation(*key, **kwargs)
435 def dynamic_operation(*key, **kwargs):
436 self.p.kwargs.update(kwargs)
--> 437 return self._process(map_obj[key], key)
438 if isinstance(self.p.operation, Operation):
439 return OperationCallable(dynamic_operation, inputs=[map_obj],
~/area51/holoviews/holoviews/core/spaces.py in __getitem__(self, key)
1130 # Not a cross product and nothing cached so compute element.
1131 if cache is not None: return cache
-> 1132 val = self._execute_callback(*tuple_key)
1133 if data_slice:
1134 val = self._dataslice(val, data_slice)
~/area51/holoviews/holoviews/core/spaces.py in _execute_callback(self, *args)
909
910 with dynamicmap_memoization(self.callback, self.streams):
--> 911 retval = self.callback(*args, **kwargs)
912 return self._style(retval)
913
~/area51/holoviews/holoviews/core/spaces.py in __call__(self, *args, **kwargs)
570
571 try:
--> 572 ret = self.callable(*args, **kwargs)
573 except KeyError:
574 # KeyError is caught separately because it is used to signal
~/area51/holoviews/holoviews/util/__init__.py in dynamic_operation(*key, **kwargs)
431 self.p.kwargs.update(kwargs)
432 obj = map_obj[key] if isinstance(map_obj, HoloMap) else map_obj
--> 433 return self._process(obj, key)
434 else:
435 def dynamic_operation(*key, **kwargs):
~/area51/holoviews/holoviews/util/__init__.py in _process(self, element, key)
417 kwargs = {k: v for k, v in self.p.kwargs.items()
418 if k in self.p.operation.params()}
--> 419 return self.p.operation.process_element(element, key, **kwargs)
420 else:
421 return self.p.operation(element, **self.p.kwargs)
~/area51/holoviews/holoviews/core/operation.py in process_element(self, element, key, **params)
141 """
142 self.p = param.ParamOverrides(self, params)
--> 143 return self._apply(element, key)
144
145
~/area51/holoviews/holoviews/core/operation.py in _apply(self, element, key)
119 for hook in self._preprocess_hooks:
120 kwargs.update(hook(self, element))
--> 121 ret = self._process(element, key)
122 for hook in self._postprocess_hooks:
123 ret = hook(self, ret, **kwargs)
~/area51/holoviews/holoviews/operation/datashader.py in _process(self, element, key)
912
913 def _process(self, element, key=None):
--> 914 agg = rasterize._process(self, element, key)
915 shaded = shade._process(self, agg, key)
916 return shaded
~/area51/holoviews/holoviews/operation/datashader.py in _process(self, element, key)
743 op = transform.instance(**op_params)
744 op._precomputed = self._precomputed
--> 745 element = element.map(op, predicate)
746 self._precomputed = op._precomputed
747 return element
~/area51/holoviews/holoviews/core/dimension.py in map(self, map_fn, specs, clone)
689 if new_val is not None:
690 deep_mapped[k] = new_val
--> 691 if applies: deep_mapped = map_fn(deep_mapped)
692 return deep_mapped
693 else:
~/area51/holoviews/holoviews/core/operation.py in __call__(self, element, **params)
161 operation=self, kwargs=params)
162 elif isinstance(element, ViewableElement):
--> 163 processed = self._apply(element)
164 elif isinstance(element, DynamicMap):
165 if any((not d.values) for d in element.kdims):
~/area51/holoviews/holoviews/core/operation.py in _apply(self, element, key)
119 for hook in self._preprocess_hooks:
120 kwargs.update(hook(self, element))
--> 121 ret = self._process(element, key)
122 for hook in self._postprocess_hooks:
123 ret = hook(self, ret, **kwargs)
~/area51/holoviews/holoviews/operation/datashader.py in _process(self, element, key)
413 ((isinstance(agg_fn, (ds.count, ds.sum, ds.mean)) and agg_fn.column not in element.kdims) or
414 (isinstance(agg_fn, ds.count_cat) and agg_fn.column in element.kdims))):
--> 415 return self._aggregate_ndoverlay(element, agg_fn)
416
417 if element._plot_id in self._precomputed:
~/area51/holoviews/holoviews/operation/datashader.py in _aggregate_ndoverlay(self, element, agg_fn)
360 grouped = element.groupby([agg_fn.column], container_type=NdOverlay,
361 group_type=NdOverlay)
--> 362 return grouped.clone({k: agg_fn1(v) for k, v in grouped.items()})
363
364 # Create aggregate instance for sum, count operations, breaking mean
~/area51/holoviews/holoviews/operation/datashader.py in <dictcomp>(.0)
360 grouped = element.groupby([agg_fn.column], container_type=NdOverlay,
361 group_type=NdOverlay)
--> 362 return grouped.clone({k: agg_fn1(v) for k, v in grouped.items()})
363
364 # Create aggregate instance for sum, count operations, breaking mean
~/area51/holoviews/holoviews/core/operation.py in __call__(self, element, **params)
161 operation=self, kwargs=params)
162 elif isinstance(element, ViewableElement):
--> 163 processed = self._apply(element)
164 elif isinstance(element, DynamicMap):
165 if any((not d.values) for d in element.kdims):
~/area51/holoviews/holoviews/core/operation.py in _apply(self, element, key)
119 for hook in self._preprocess_hooks:
120 kwargs.update(hook(self, element))
--> 121 ret = self._process(element, key)
122 for hook in self._postprocess_hooks:
123 ret = hook(self, ret, **kwargs)
~/area51/holoviews/holoviews/operation/datashader.py in _process(self, element, key)
450
451 dfdata = PandasInterface.as_dframe(data)
--> 452 agg = getattr(cvs, glyph)(dfdata, x.name, y.name, agg_fn)
453 if 'x_axis' in agg.coords and 'y_axis' in agg.coords:
454 agg = agg.rename({'x_axis': x, 'y_axis': y})
~/jungle/local/lib/python3.5/site-packages/datashader/core.py in line(self, source, x, y, agg)
185 if agg is None:
186 agg = any_rdn()
--> 187 return bypixel(source, self, Line(x, y), agg)
188
189 # TODO re 'untested', below: Consider replacing with e.g. a 3x3
~/jungle/local/lib/python3.5/site-packages/datashader/core.py in bypixel(source, canvas, glyph, agg)
535 agg.validate(schema)
536 canvas.validate()
--> 537 return bypixel.pipeline(source, schema, canvas, glyph, agg)
538
539
~/jungle/local/lib/python3.5/site-packages/datashader/utils.py in __call__(self, head, *rest, **kwargs)
56 typ = type(head)
57 if typ in lk:
---> 58 return lk[typ](head, *rest, **kwargs)
59 for cls in getmro(typ)[1:]:
60 if cls in lk:
~/jungle/local/lib/python3.5/site-packages/datashader/dask.py in dask_pipeline(df, schema, canvas, glyph, summary)
19 dsk, name = glyph_dispatch(glyph, df, schema, canvas, summary)
20
---> 21 get = _globals['get'] or getattr(df, '__dask_scheduler__', None) or df._default_get
22 keys = getattr(df, '__dask_keys__', None) or df._keys
23 optimize = getattr(df, '__dask_optimize__', None) or df._optimize
KeyError: 'get'
I tried doing a loc version by overlaying plots with data locced by category, with plain Holoviews,
and it throws an error:
~/jungle/local/lib/python3.5/site-packages/ipython-6.3.1-py3.5.egg/IPython/core/formatters.py in __call__(self, obj, include, exclude)
968
969 if method is not None:
--> 970 return method(include=include, exclude=exclude)
971 return None
972 else:
~/area51/holoviews/holoviews/core/dimension.py in _repr_mimebundle_(self, include, exclude)
1229 combined and returned.
1230 """
-> 1231 return Store.render(self)
1232
1233
~/area51/holoviews/holoviews/core/options.py in render(cls, obj)
1287 data, metadata = {}, {}
1288 for hook in hooks:
-> 1289 ret = hook(obj)
1290 if ret is None:
1291 continue
~/area51/holoviews/holoviews/ipython/display_hooks.py in pprint_display(obj)
270 if not ip.display_formatter.formatters['text/plain'].pprint:
271 return None
--> 272 return display(obj, raw_output=True)
273
274
~/area51/holoviews/holoviews/ipython/display_hooks.py in display(obj, raw_output, **kwargs)
246 elif isinstance(obj, (HoloMap, DynamicMap)):
247 with option_state(obj):
--> 248 output = map_display(obj)
249 elif isinstance(obj, Plot):
250 output = render(obj)
~/area51/holoviews/holoviews/ipython/display_hooks.py in wrapped(element)
140 try:
141 max_frames = OutputSettings.options['max_frames']
--> 142 mimebundle = fn(element, max_frames=max_frames)
143 if mimebundle is None:
144 return {}, {}
~/area51/holoviews/holoviews/ipython/display_hooks.py in map_display(vmap, max_frames)
200 return None
201
--> 202 return render(vmap)
203
204
~/area51/holoviews/holoviews/ipython/display_hooks.py in render(obj, **kwargs)
63 renderer = renderer.instance(fig='png')
64
---> 65 return renderer.components(obj, **kwargs)
66
67
~/area51/holoviews/holoviews/plotting/bokeh/renderer.py in components(self, obj, fmt, comm, **kwargs)
260 # Bokeh has to handle comms directly in <0.12.15
261 comm = False if bokeh_version < '0.12.15' else comm
--> 262 return super(BokehRenderer, self).components(obj,fmt, comm, **kwargs)
263
264
~/area51/holoviews/holoviews/plotting/renderer.py in components(self, obj, fmt, comm, **kwargs)
327 plot = obj
328 else:
--> 329 plot, fmt = self._validate(obj, fmt)
330
331 widget_id = None
~/area51/holoviews/holoviews/plotting/renderer.py in _validate(self, obj, fmt, **kwargs)
226 if isinstance(obj, tuple(self.widgets.values())):
227 return obj, 'html'
--> 228 plot = self.get_plot(obj, renderer=self, **kwargs)
229
230 fig_formats = self.mode_formats['fig'][self.mode]
~/area51/holoviews/holoviews/plotting/bokeh/renderer.py in get_plot(self_or_cls, obj, doc, renderer)
153 curdoc().theme = self_or_cls.theme
154 doc.theme = self_or_cls.theme
--> 155 plot = super(BokehRenderer, self_or_cls).get_plot(obj, renderer)
156 plot.document = doc
157 return plot
~/area51/holoviews/holoviews/plotting/renderer.py in get_plot(self_or_cls, obj, renderer)
209 plot_opts = self_or_cls.plot_options(obj, self_or_cls.size)
210 plot = self_or_cls.plotting_class(obj)(obj, renderer=renderer,
--> 211 **plot_opts)
212 defaults = [kd.default for kd in plot.dimensions]
213 init_key = tuple(v if d is None else d for v, d in
~/area51/holoviews/holoviews/plotting/plot.py in __init__(self, overlay, ranges, batched, keys, group_counter, **params)
889 self.group_counter = Counter() if group_counter is None else group_counter
890 self.zoffset = 0
--> 891 self.subplots = self._create_subplots(ranges)
892 self.traverse(lambda x: setattr(x, 'comm', self.comm))
893 self.top_level = keys is None
~/area51/holoviews/holoviews/plotting/plot.py in _create_subplots(self, ranges)
939 if isinstance(self.hmap, DynamicMap):
940 dmap_streams = [get_nested_streams(layer) for layer in
--> 941 split_dmap_overlay(self.hmap)]
942 else:
943 dmap_streams = [None]*len(keys)
~/area51/holoviews/holoviews/plotting/util.py in split_dmap_overlay(obj, depth)
217 if obj.callback.inputs and is_dynamic_overlay(obj):
218 for inp in obj.callback.inputs:
--> 219 layers += split_dmap_overlay(inp, depth+1)
220 else:
221 for v in obj.last.values():
~/area51/holoviews/holoviews/plotting/util.py in split_dmap_overlay(obj, depth)
217 if obj.callback.inputs and is_dynamic_overlay(obj):
218 for inp in obj.callback.inputs:
--> 219 layers += split_dmap_overlay(inp, depth+1)
220 else:
221 for v in obj.last.values():
~/area51/holoviews/holoviews/plotting/util.py in split_dmap_overlay(obj, depth)
217 if obj.callback.inputs and is_dynamic_overlay(obj):
218 for inp in obj.callback.inputs:
--> 219 layers += split_dmap_overlay(inp, depth+1)
220 else:
221 for v in obj.last.values():
~/area51/holoviews/holoviews/plotting/util.py in split_dmap_overlay(obj, depth)
211 layers = []
212 if isinstance(obj, DynamicMap):
--> 213 if issubclass(obj.type, NdOverlay) and not depth:
214 for v in obj.last.values():
215 layers.append(obj)
TypeError: issubclass() arg 1 must be a class
I am confused about how quadmesh represents data and coordinates. In my head, the coordinates of a quadmesh specify the positions of cell vertices, while the data represent values for the entire cell. Since there are more vertices than there are cells, the size of the coordinates should be greater. Specifically, if the shape of the data is (M, N), the x and y coordinates for the quadmesh should both have the shape (M+1, N+1).
This seems to be at odds with the quadmesh example, in which the coordinates and data have the exact same shape:
n = 20
coords = np.linspace(-1.5,1.5,n)
X,Y = np.meshgrid(coords, coords);
Qx = np.cos(Y) - np.cos(X)
Qy = np.sin(Y) + np.sin(X)
Z = np.sqrt(X**2 + Y**2)
print('Shape of x-coordinates:', Qx.shape)
print('Shape of y-coordinates:', Qy.shape)
print('Shape of value array:', Z.shape)
qmesh = hv.QuadMesh((Qx, Qy, Z))
qmeshShape of x-coordinates: (20, 20)
Shape of y-coordinates: (20, 20)
Shape of value array: (20, 20)
However, I find that I can in fact do
Z = np.sqrt(X**2 + Y**2)[:-1, :-1]with no discernible difference in output. This fits my mental model much better.
What is going on here?
A declarative, efficient, and flexible JavaScript library for building user interfaces.
🖖 Vue.js is a progressive, incrementally-adoptable JavaScript framework for building UI on the web.
TypeScript is a superset of JavaScript that compiles to clean JavaScript output.
An Open Source Machine Learning Framework for Everyone
The Web framework for perfectionists with deadlines.
A PHP framework for web artisans
Bring data to life with SVG, Canvas and HTML. 📊📈🎉
JavaScript (JS) is a lightweight interpreted programming language with first-class functions.
Some thing interesting about web. New door for the world.
A server is a program made to process requests and deliver data to clients.
Machine learning is a way of modeling and interpreting data that allows a piece of software to respond intelligently.
Some thing interesting about visualization, use data art
Some thing interesting about game, make everyone happy.
We are working to build community through open source technology. NB: members must have two-factor auth.
Open source projects and samples from Microsoft.
Google ❤️ Open Source for everyone.
Alibaba Open Source for everyone
Data-Driven Documents codes.
China tencent open source team.